Histone_ChIP-seq_ENCSR000ASD
| Id: | Histone_ChIP-seq/ENCSR000ASD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ASD [biosamplesummary="Homo sapiens endothelial cell of umbilical vein newborn" and target="H3K79me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: newborn output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN973APA|/analyses/ENCAN973APA/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF251MRD|/files/ENCFF251MRD/} processed by ChIP-seq ENCODE3 hg19 pipeline has 15488849 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me2-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF561ROR|/files/ENCFF561ROR/} processed by ChIP-seq ENCODE3 hg19 pipeline has 28803309 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K79me2-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
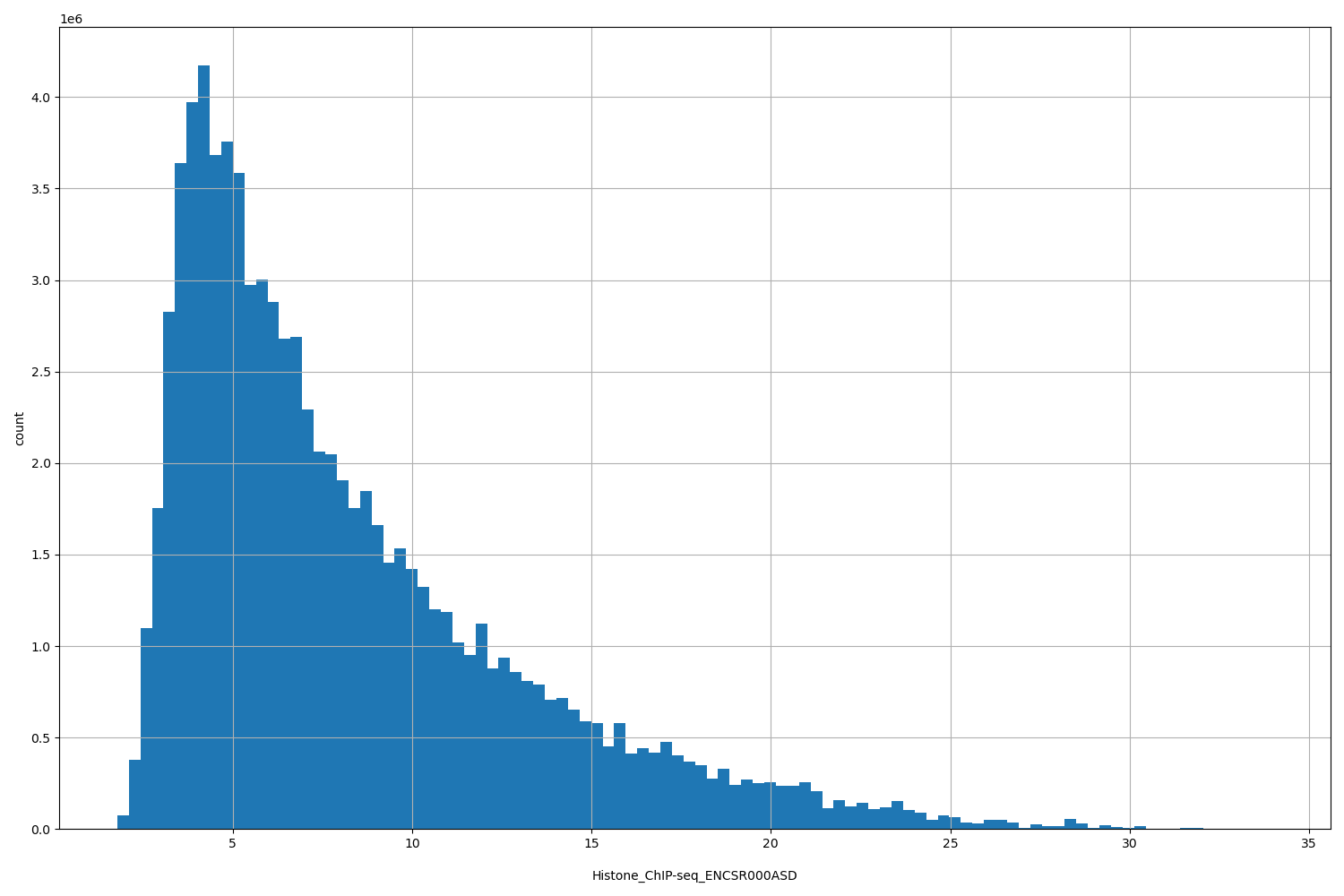
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ASD | float |
Histone_ChIP-seq_ENCSR000ASD |
Histone_ChIP-seq ENCSR000ASD [biosample_summary="Homo sapiens endothelial cell of umbilical vein newborn" and target="H3K79me2"]
|
 |
[1.79, 34] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF229RMW.bed.gz | 2.85 MB | c8ae6ddc028fd4f1bed7f7b08aef3f8e |
| ENCFF229RMW.bed.gz.dvc | 101.0 B | 4fa8bfbacd46d4f7edc981dd0588280f |
| ENCFF229RMW.tabix.bed.gz | 1.36 MB | 1a645d3756aa287dbd4a46e54f4f2850 |
| ENCFF229RMW.tabix.bed.gz.dvc | 107.0 B | e25d22eff453431d9dae46b1849f9218 |
| ENCFF229RMW.tabix.bed.gz.tbi | 132.0 KB | 9a9ec504b718dc0bd64af7c43d220d62 |
| ENCFF229RMW.tabix.bed.gz.tbi.dvc | 110.0 B | 8099cc57a31d628a765da018aaee6c36 |
| genomic_resource.yaml | 2.61 KB | d7d595ef0e5e722e032746e8bcbbd324 |
| genomic_resource_original.yaml | 2.47 KB | baa8fcbeebe035af4ae360e5991d3576 |
| statistics/ |