Histone_ChIP-seq_ENCSR000ARN
| Id: | Histone_ChIP-seq/ENCSR000ARN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ARN [biosamplesummary="Homo sapiens keratinocyte female" and target="H3K9me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN688YHT|/analyses/ENCAN688YHT/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN688YHT|/analyses/ENCAN688YHT/} has in progress subobject document {264329f7-a171-41d7-9bbb-e9cc74ad83ab|/documents/264329f7-a171-41d7-9bbb-e9cc74ad83ab/} audit_warning: Processed unfiltered alignments file {ENCFF049TZA|/files/ENCFF049TZA/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 38545632 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed unfiltered alignments file {ENCFF460ZZP|/files/ENCFF460ZZP/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 40473616 mapped reads. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K9me3-human and investigated as a broad histone mark is 35 million mapped reads. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
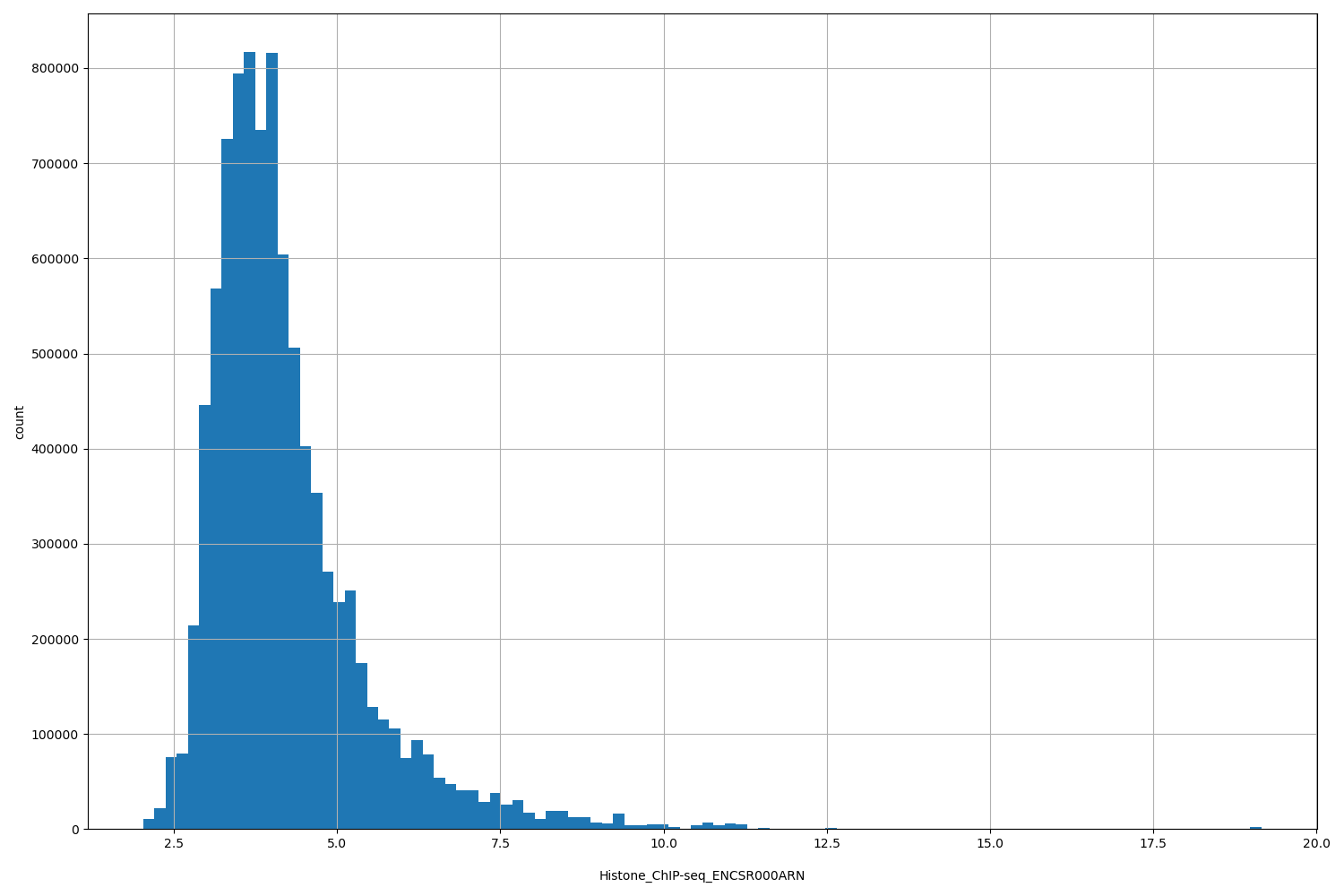
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ARN | float |
Histone_ChIP-seq_ENCSR000ARN |
Histone_ChIP-seq ENCSR000ARN [biosample_summary="Homo sapiens keratinocyte female" and target="H3K9me3"]
|
 |
[2.03, 19.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF403AZM.bed.gz | 524.38 KB | 01ecb34899ef175d51bf23830a259999 |
| ENCFF403AZM.bed.gz.dvc | 100.0 B | 1873aca12f3b472a4aece23bdc807c74 |
| ENCFF403AZM.tabix.bed.gz | 289.49 KB | 7398e18e6deef0adf8a45ec91d337b07 |
| ENCFF403AZM.tabix.bed.gz.dvc | 106.0 B | 977bd5ddc7f3592006f42999c631583c |
| ENCFF403AZM.tabix.bed.gz.tbi | 140.4 KB | 64446a8d8123f3ed39076cf52e4f6d15 |
| ENCFF403AZM.tabix.bed.gz.tbi.dvc | 110.0 B | 1929a93ba25c0d84c2098b68090de2f9 |
| genomic_resource.yaml | 2.6 KB | 1b274ef1af41db67f050ce7e407699d5 |
| genomic_resource_original.yaml | 2.48 KB | 8d88833cbbb573e717085ac1815f14d7 |
| statistics/ |