Histone_ChIP-seq_ENCSR000ARF
| Id: | Histone_ChIP-seq/ENCSR000ARF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ARF [biosamplesummary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="H2AFZ"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female adult (50 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN601EVX|/analyses/ENCAN601EVX/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN601EVX|/analyses/ENCAN601EVX/} has in progress subobject document {903b88ff-6c6a-4b12-8bd5-53ce947f60da|/documents/903b88ff-6c6a-4b12-8bd5-53ce947f60da/} audit_warning: Processed alignments file {ENCFF338WQQ|/files/ENCFF338WQQ/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 17045512 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2AFZ-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF940IDG|/files/ENCFF940IDG/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 14833881 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H2AFZ-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
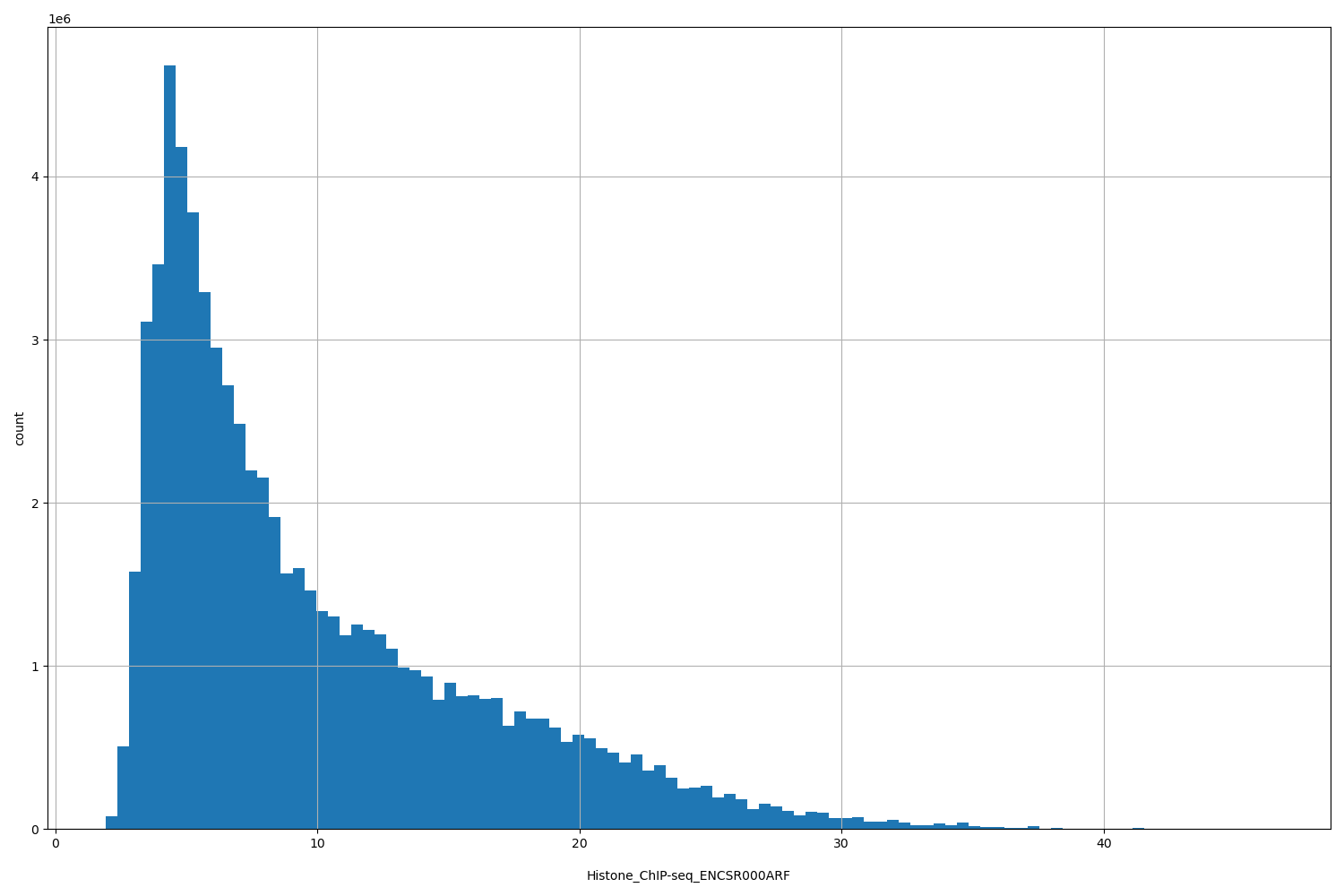
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ARF | float |
Histone_ChIP-seq_ENCSR000ARF |
Histone_ChIP-seq ENCSR000ARF [biosample_summary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="H2AFZ"]
|
 |
[1.94, 46.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF360HTG.bed.gz | 2.44 MB | 9e79ae59a6489ad4e8bf960995552c0b |
| ENCFF360HTG.bed.gz.dvc | 101.0 B | 424b0631e1528ffa00254e6391b71c33 |
| ENCFF360HTG.tabix.bed.gz | 1.05 MB | 6c672c8f7597cd5bb1f3e32e2eb66d5b |
| ENCFF360HTG.tabix.bed.gz.dvc | 107.0 B | 994a1041165bbb0340035f949aff07e1 |
| ENCFF360HTG.tabix.bed.gz.tbi | 375.28 KB | 3404ec8b1034b525ebd498e1137c8f17 |
| ENCFF360HTG.tabix.bed.gz.tbi.dvc | 110.0 B | a71579660f99574100d9ffb07728f1c2 |
| genomic_resource.yaml | 2.7 KB | 2eb2a23a24d72d2ab602ce2775a1d995 |
| genomic_resource_original.yaml | 2.56 KB | 0face60757386d8a196338548a95af90 |
| statistics/ |