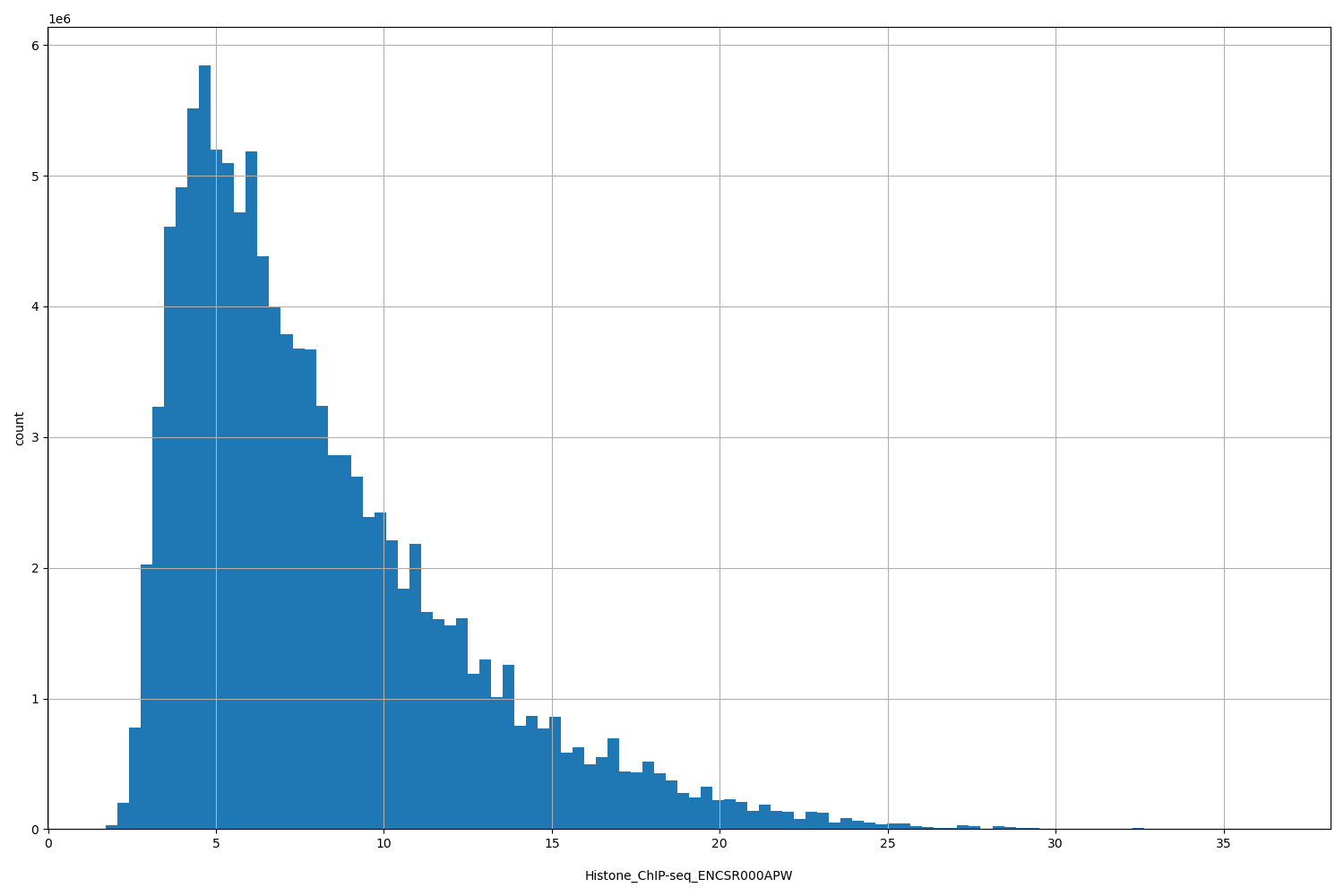
Histone_ChIP-seq_ENCSR000APW
| Id: | Histone_ChIP-seq/ENCSR000APW |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000APW [biosamplesummary="Homo sapiens HeLa-S3" and target="H3K4me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN319FKG|/analyses/ENCAN319FKG/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF617YCQ|/files/ENCFF617YCQ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 15505318 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF235FMM|/files/ENCFF235FMM/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18772478 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000APW | float |
Histone_ChIP-seq_ENCSR000APW |
Histone_ChIP-seq ENCSR000APW [biosample_summary="Homo sapiens HeLa-S3" and target="H3K4me1"]
|
 |
[1.73, 36.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF360CQR.bed.gz | 2.49 MB | 102441e09b1b270ceffcfea1f9103951 |
| ENCFF360CQR.bed.gz.dvc | 101.0 B | 14f84bead03a8b80e8066e24bc99956d |
| ENCFF360CQR.tabix.bed.gz | 1.14 MB | 042af90ba1d5e452f089fafee2352299 |
| ENCFF360CQR.tabix.bed.gz.dvc | 107.0 B | 0688ad7ad10b26e8b1fee3957c8a9b09 |
| ENCFF360CQR.tabix.bed.gz.tbi | 299.32 KB | 4baab26679bf7636d5c65aec6821ef0c |
| ENCFF360CQR.tabix.bed.gz.tbi.dvc | 110.0 B | 90f55d6c25a0391df49347a786c5279f |
| genomic_resource.yaml | 2.37 KB | 513dce86770ca9e3b46b6599127a222d |
| genomic_resource_original.yaml | 2.26 KB | 382438abb12a470032f1514d6989d6c2 |
| statistics/ |