Histone_ChIP-seq_ENCSR000APR
| Id: | Histone_ChIP-seq/ENCSR000APR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000APR [biosamplesummary="Homo sapiens fibroblast of dermis NONE and female adult" and target="H3K4me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female adult output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN172DGC|/analyses/ENCAN172DGC/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF664FYA|/files/ENCFF664FYA/} processed by ChIP-seq ENCODE3 hg19 pipeline has 9116188 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me3-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
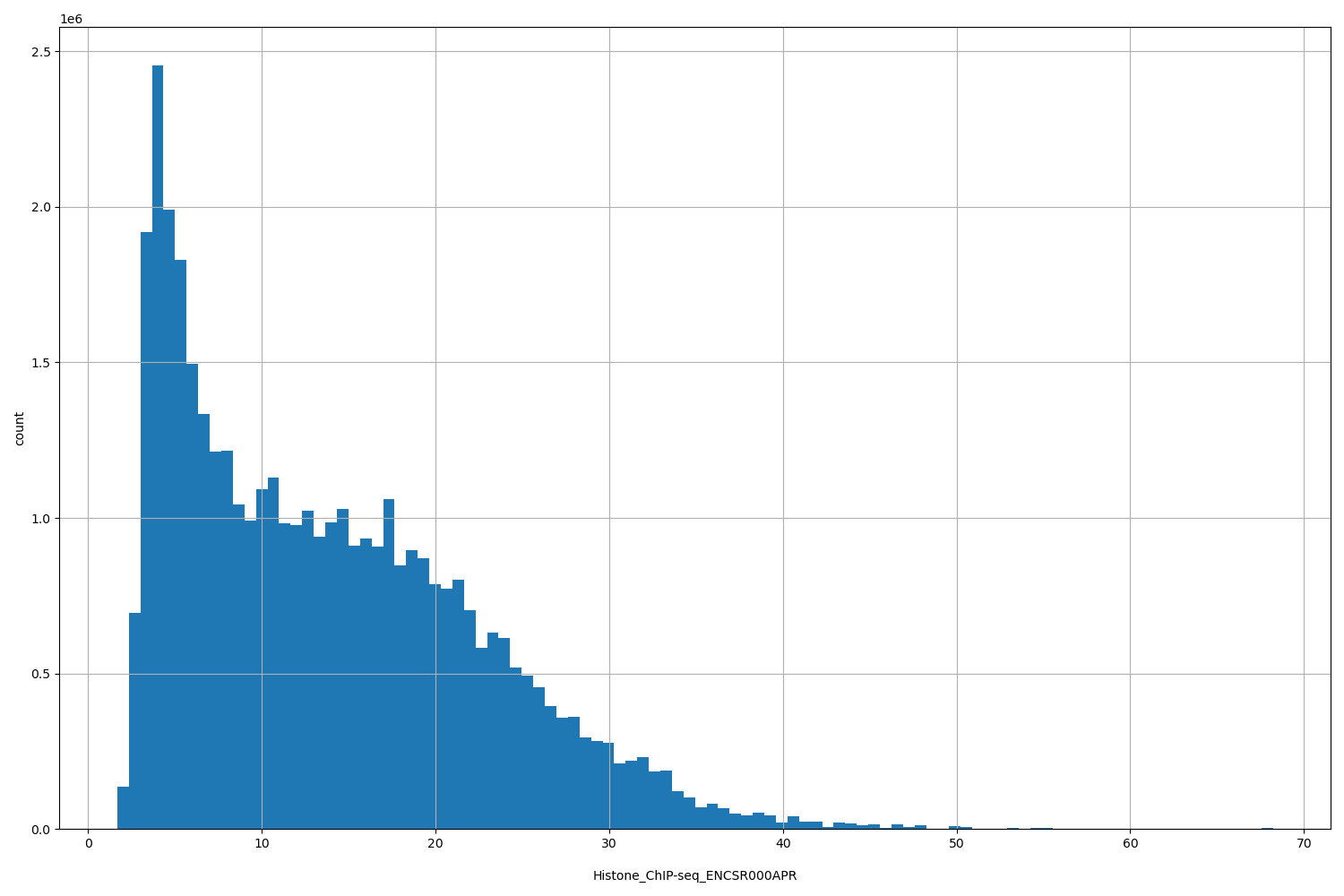
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000APR | float |
Histone_ChIP-seq_ENCSR000APR |
Histone_ChIP-seq ENCSR000APR [biosample_summary="Homo sapiens fibroblast of dermis NONE and female adult" and target="H3K4me3"]
|
 |
[1.67, 68.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF811YTI.bed.gz | 1.23 MB | 822e2f42afa58c13f6e1184e43a2c061 |
| ENCFF811YTI.bed.gz.dvc | 101.0 B | c1d1303198def10744993efa03c0f23a |
| ENCFF811YTI.tabix.bed.gz | 543.57 KB | f24ef8e8d0d055bb5ce3992b0a4a4ac1 |
| ENCFF811YTI.tabix.bed.gz.dvc | 106.0 B | f4c8df024499251149f71ac68ea1a992 |
| ENCFF811YTI.tabix.bed.gz.tbi | 217.2 KB | 9799458a23664138a2eceaf0300905a2 |
| ENCFF811YTI.tabix.bed.gz.tbi.dvc | 110.0 B | 3518c17808268924542200e5757eba1a |
| genomic_resource.yaml | 1.97 KB | a5b75b8769eb3f3237f887763c531546 |
| genomic_resource_original.yaml | 1.83 KB | 63804b278a55dce04eec083aaeb3919c |
| statistics/ |