Histone_ChIP-seq_ENCSR000AOQ
| Id: | Histone_ChIP-seq/ENCSR000AOQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AOQ [biosamplesummary="Homo sapiens astrocyte" and target="H3K27ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN749XVU|/analyses/ENCAN749XVU/} has in progress subobject document {bf944498-04a0-44a2-9185-e6dc2cde2841|/documents/bf944498-04a0-44a2-9185-e6dc2cde2841/} audit_internal_action: Released analysis {ENCAN749XVU|/analyses/ENCAN749XVU/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_warning: Processed alignments file {ENCFF759CXE|/files/ENCFF759CXE/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 10226128 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF312ZIP|/files/ENCFF312ZIP/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 10925509 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
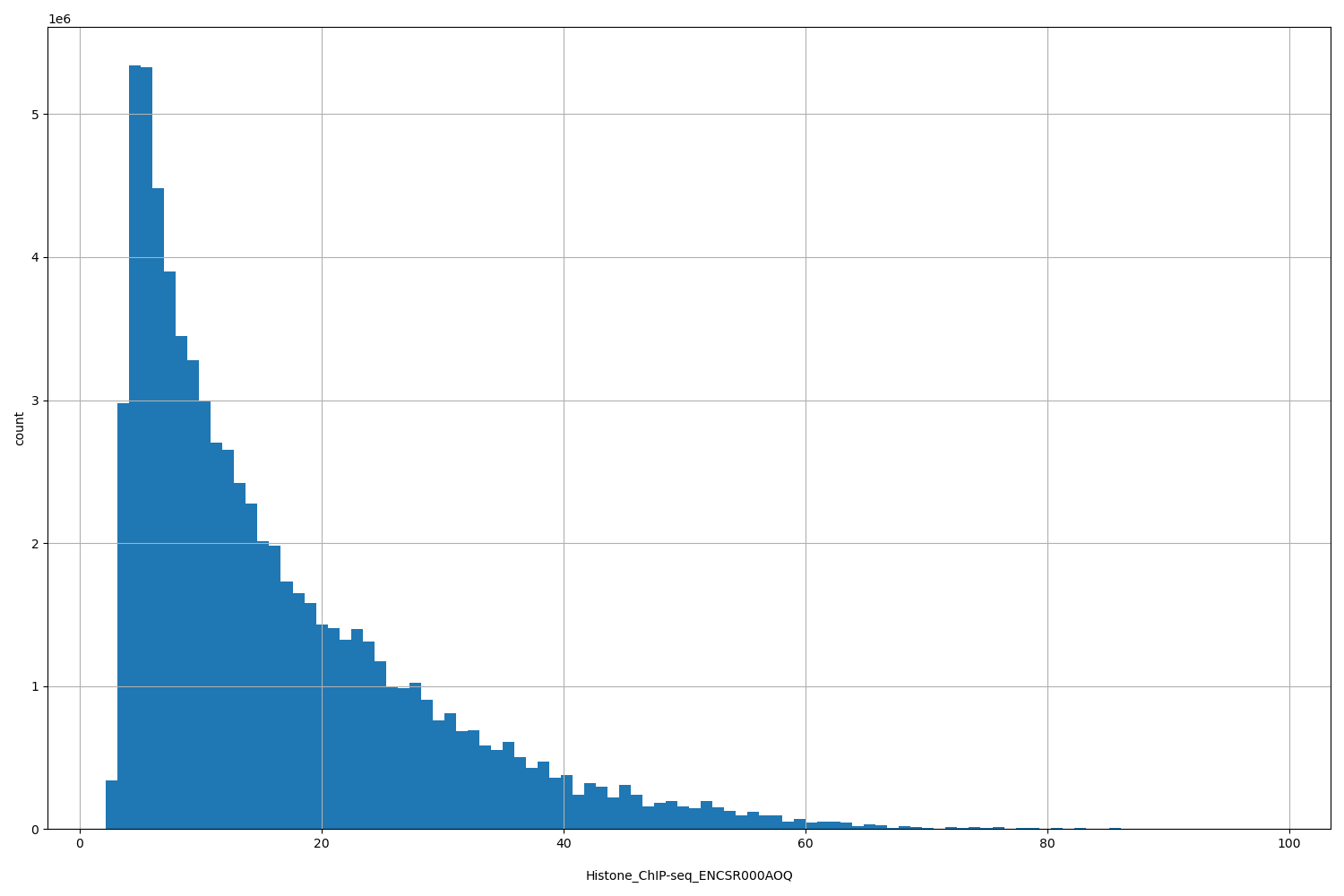
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AOQ | float |
Histone_ChIP-seq_ENCSR000AOQ |
Histone_ChIP-seq ENCSR000AOQ [biosample_summary="Homo sapiens astrocyte" and target="H3K27ac"]
|
 |
[2.18, 98.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF138TME.bed.gz | 1.86 MB | ca601df322d63ea3c1d2b560a1d5dbc4 |
| ENCFF138TME.bed.gz.dvc | 101.0 B | 71ac94f2b401aca0c460132bdcaef767 |
| ENCFF138TME.tabix.bed.gz | 807.35 KB | b83aa2131d9d2607ad5c3310b3813bbf |
| ENCFF138TME.tabix.bed.gz.dvc | 106.0 B | 177358852fd96bb88fa05172cfebd05e |
| ENCFF138TME.tabix.bed.gz.tbi | 269.62 KB | 158a4302999de7acab279952d75d3463 |
| ENCFF138TME.tabix.bed.gz.tbi.dvc | 110.0 B | 5eedb48c6e4a9720c8f5beb206661b2d |
| genomic_resource.yaml | 2.61 KB | e555f9de5ae4b03731099dea28c70384 |
| genomic_resource_original.yaml | 2.5 KB | e0b6cb3e55e3c61f610d3eefba621a9c |
| statistics/ |