Histone_ChIP-seq_ENCSR000ANJ
| Id: | Histone_ChIP-seq/ENCSR000ANJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ANJ [biosamplesummary="Homo sapiens skeletal muscle myoblast male adult (22 years)" and target="H3K4me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (22 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN253XGW|/analyses/ENCAN253XGW/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN253XGW|/analyses/ENCAN253XGW/} has in progress subobject document {3e2dc527-c37d-4c1f-a60d-3ec14913ffe6|/documents/3e2dc527-c37d-4c1f-a60d-3ec14913ffe6/} audit_not_compliant: Processed alignments file {ENCFF976IIM|/files/ENCFF976IIM/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 6434801 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF764HBL|/files/ENCFF764HBL/} processed by ChIP-seq ENCODE4 v1.6.0 GRCh38 pipeline has 17868990 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
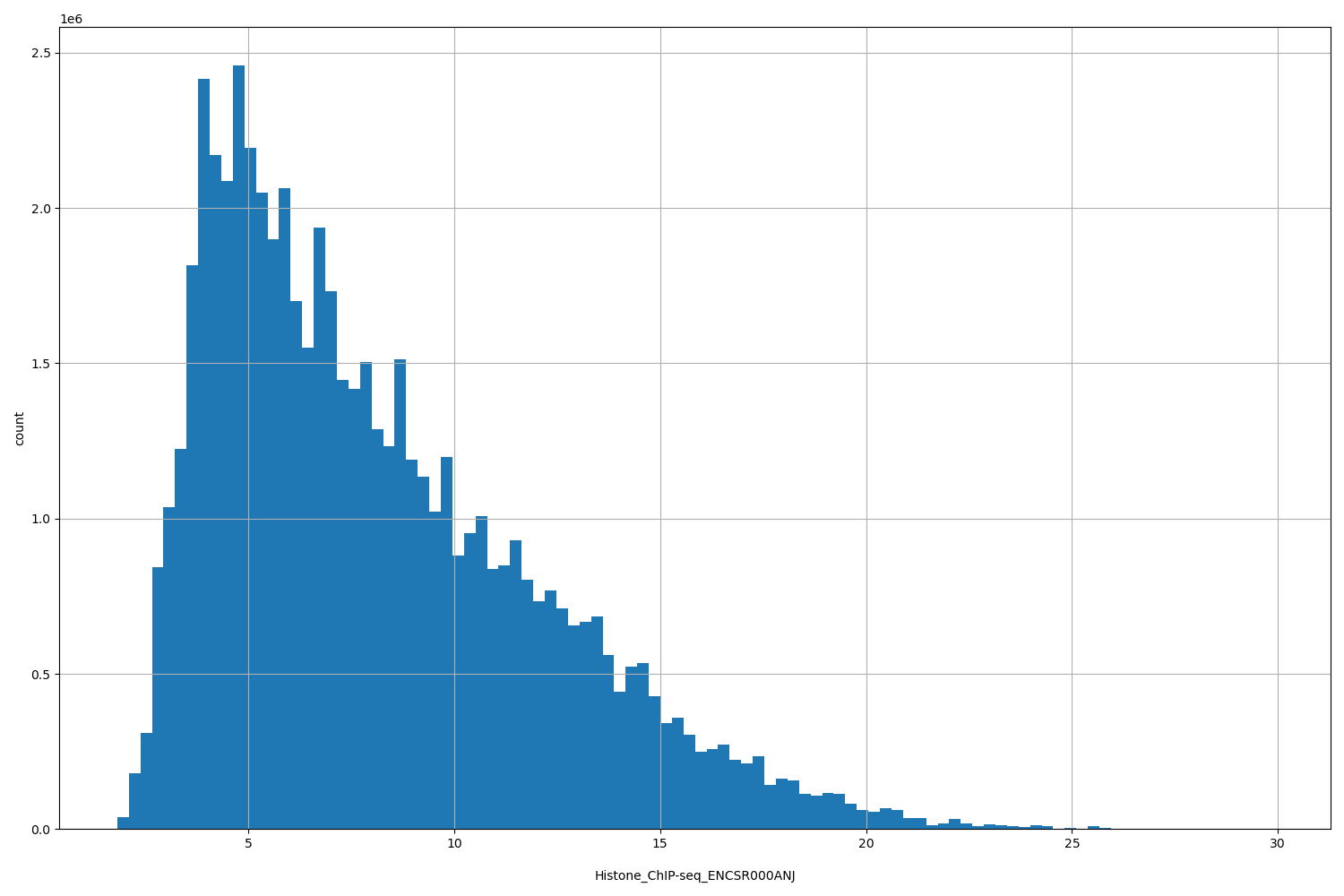
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ANJ | float |
Histone_ChIP-seq_ENCSR000ANJ |
Histone_ChIP-seq ENCSR000ANJ [biosample_summary="Homo sapiens skeletal muscle myoblast male adult (22 years)" and target="H3K4me2"]
|
 |
[1.82, 29.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF554YVU.bed.gz | 2.03 MB | a2fa7578f7c738dca53a1a9f67b45a7b |
| ENCFF554YVU.bed.gz.dvc | 101.0 B | 3ba2ba50f9c70dd2a2974c9c8fb305b7 |
| ENCFF554YVU.tabix.bed.gz | 970.33 KB | fa01821fe94d662ba3f542e039c55da7 |
| ENCFF554YVU.tabix.bed.gz.dvc | 106.0 B | 10c5639f33ecdbebef62532f6ea2700b |
| ENCFF554YVU.tabix.bed.gz.tbi | 320.62 KB | 8c44e9c70221064320c24b6334cb3f8c |
| ENCFF554YVU.tabix.bed.gz.tbi.dvc | 110.0 B | 76144d0143841ed3b673c8725ce4ad92 |
| genomic_resource.yaml | 2.71 KB | 4712ed7bb6b1a8581bec306b43e98ca5 |
| genomic_resource_original.yaml | 2.56 KB | 72cbd3b41a993054e6860870333885d0 |
| statistics/ |