Histone_ChIP-seq_ENCSR000ANC
| Id: | Histone_ChIP-seq/ENCSR000ANC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000ANC [biosamplesummary="Homo sapiens H1" and target="H3K4me2"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN890AIH|/analyses/ENCAN890AIH/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_warning: Processed alignments file {ENCFF694EXK|/files/ENCFF694EXK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 11330199 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_warning: Processed alignments file {ENCFF115ZGZ|/files/ENCFF115ZGZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 12550300 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
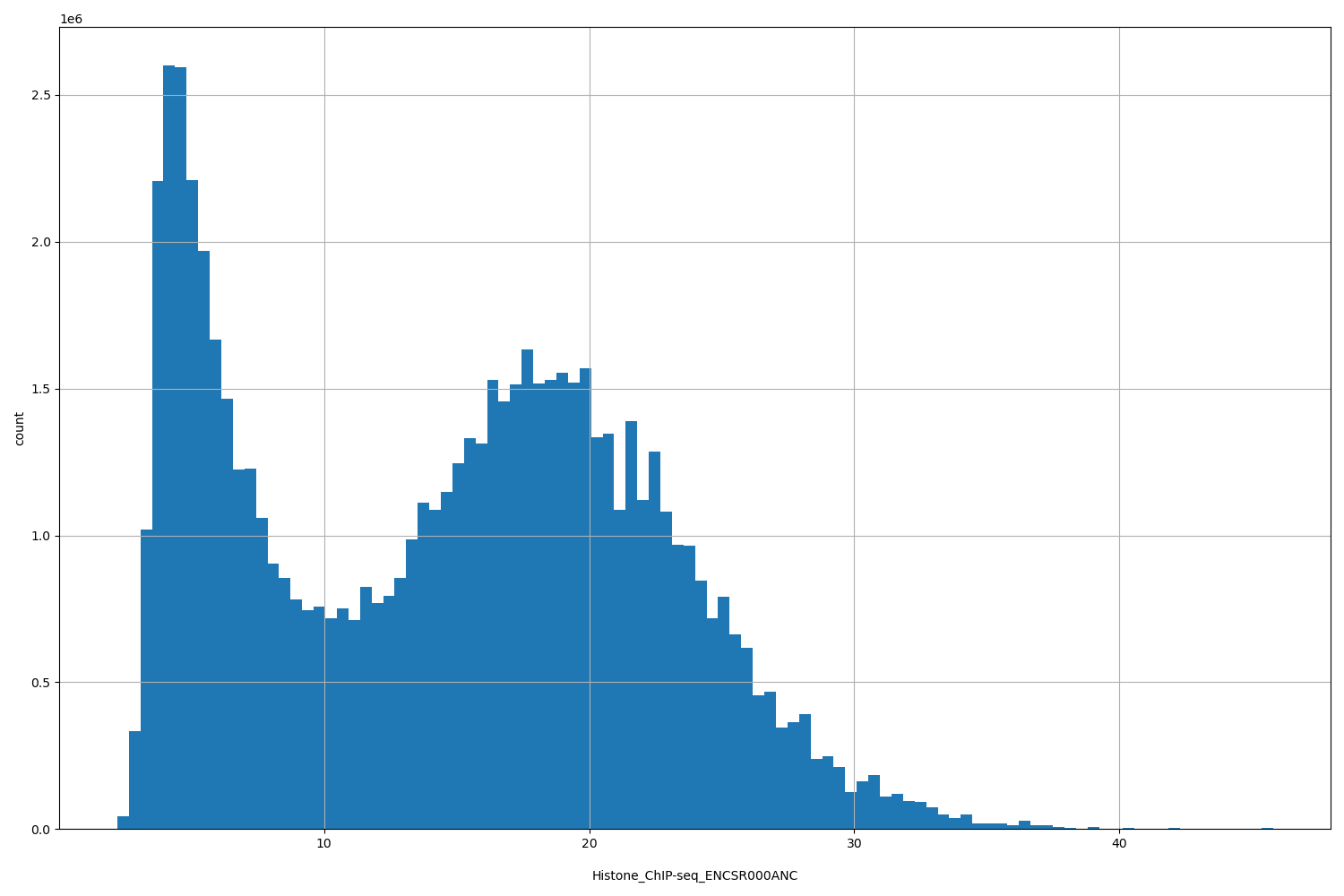
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000ANC | float |
Histone_ChIP-seq_ENCSR000ANC |
Histone_ChIP-seq ENCSR000ANC [biosample_summary="Homo sapiens H1" and target="H3K4me2"]
|
 |
[2.19, 45.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF836LZM.bed.gz | 1.78 MB | e9b56f3dfeca3718e33f56eaf04a0212 |
| ENCFF836LZM.bed.gz.dvc | 101.0 B | 377f418cd4e06325d237b2f1a05bf49f |
| ENCFF836LZM.tabix.bed.gz | 781.04 KB | fef127ad28e79f829ea3009e043edeb0 |
| ENCFF836LZM.tabix.bed.gz.dvc | 106.0 B | 73f3ee24752ee4e494b9509495614d39 |
| ENCFF836LZM.tabix.bed.gz.tbi | 298.07 KB | 91a141c65d5de14f818639b5ff4a9e32 |
| ENCFF836LZM.tabix.bed.gz.tbi.dvc | 110.0 B | 9c207f05dda78ae2a8b0d48dc0914b0e |
| genomic_resource.yaml | 2.34 KB | 2c6c61f27f17c9d6c21f4569f97afb00 |
| genomic_resource_original.yaml | 2.24 KB | 3d9a958a8860cc30355ea0dd2082a734 |
| statistics/ |