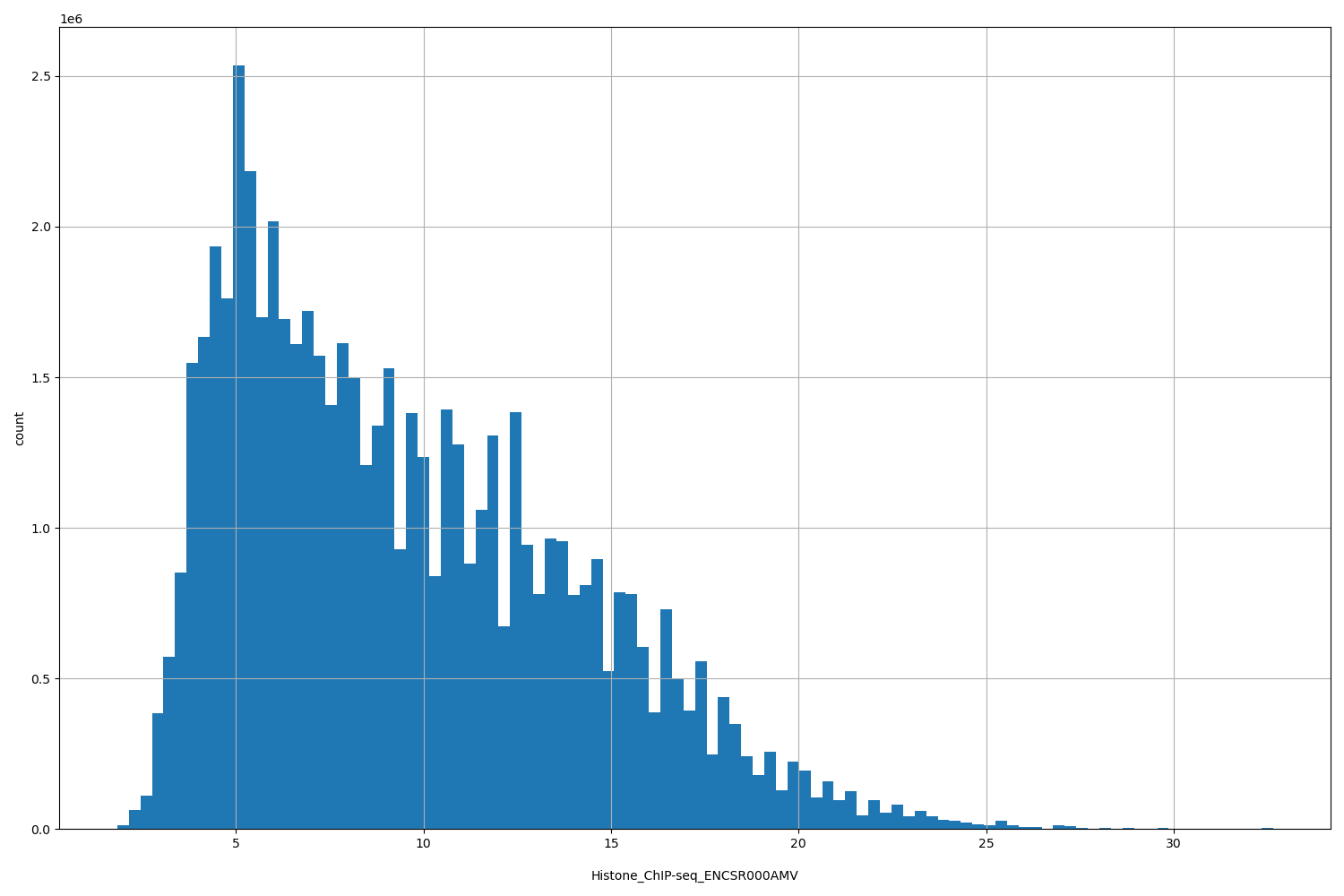
Histone_ChIP-seq_ENCSR000AMV
| Id: | Histone_ChIP-seq/ENCSR000AMV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AMV [biosamplesummary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K4me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (45 years) female child (11 years) output_type: replicated peaks audit_error: Processed alignments file {ENCFF227EHJ|/files/ENCFF227EHJ/} processed by ChIP-seq ENCODE3 hg19 pipeline has 3169809 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_internal_action: Archived analysis {ENCAN746THU|/analyses/ENCAN746THU/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF376UFR|/files/ENCFF376UFR/} processed by ChIP-seq ENCODE3 hg19 pipeline has 7784126 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AMV | float |
Histone_ChIP-seq_ENCSR000AMV |
Histone_ChIP-seq ENCSR000AMV [biosample_summary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K4me2"]
|
 |
[1.84, 32.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF862CMS.bed.gz | 1.83 MB | 2a4621b4775e155fa27fd57154eaf8b1 |
| ENCFF862CMS.bed.gz.dvc | 101.0 B | 9c495d5b8cdfb530d7cdf4a48a266111 |
| ENCFF862CMS.tabix.bed.gz | 909.6 KB | 58ebb932a03fd9abc7fd6d290e978aab |
| ENCFF862CMS.tabix.bed.gz.dvc | 106.0 B | af5ce14a24645107e1b4228349e71279 |
| ENCFF862CMS.tabix.bed.gz.tbi | 301.69 KB | 9ea8fda1aa45e53be9d88945d052174d |
| ENCFF862CMS.tabix.bed.gz.tbi.dvc | 110.0 B | 532db61644565c769cacbfcaf59b8261 |
| genomic_resource.yaml | 2.59 KB | 2d24b5ec83903a73f62bbfdb6b9a5fc3 |
| genomic_resource_original.yaml | 2.42 KB | ea55a69d7cbe256883db090116f2e292 |
| statistics/ |