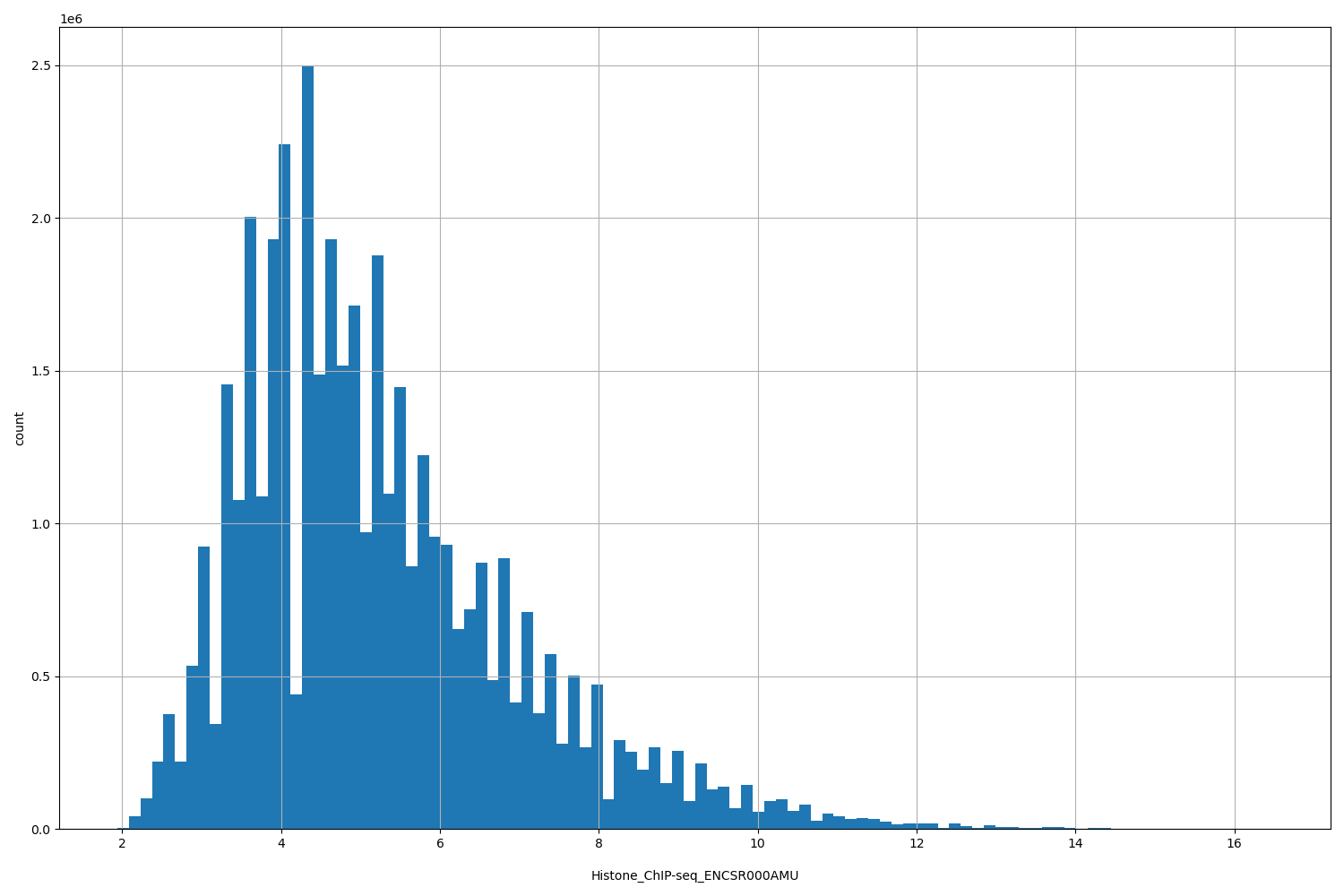
Histone_ChIP-seq_ENCSR000AMU
| Id: | Histone_ChIP-seq/ENCSR000AMU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AMU [biosamplesummary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K4me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (45 years) female child (11 years) output_type: pseudoreplicated peaks audit_error: Processed alignments file {ENCFF088FFQ|/files/ENCFF088FFQ/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 4430164 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_internal_action: Released analysis {ENCAN573OCO|/analyses/ENCAN573OCO/} has in progress subobject document {e25e2309-1110-4f34-b09e-2ee6fea83131|/documents/e25e2309-1110-4f34-b09e-2ee6fea83131/} audit_internal_action: Released analysis {ENCAN573OCO|/analyses/ENCAN573OCO/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF505WNW|/files/ENCFF505WNW/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 13259702 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AMU | float |
Histone_ChIP-seq_ENCSR000AMU |
Histone_ChIP-seq ENCSR000AMU [biosample_summary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K4me1"]
|
 |
[1.94, 16.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF928WDI.bed.gz | 1.87 MB | 191e85afc7bfaf4fdb3f396498dc7328 |
| ENCFF928WDI.bed.gz.dvc | 101.0 B | 9cfdb2924dd8384daa9522e354ff68df |
| ENCFF928WDI.tabix.bed.gz | 987.66 KB | 559c3551bd4562ea86120a1a3a47d913 |
| ENCFF928WDI.tabix.bed.gz.dvc | 107.0 B | 7474e12cf136cf9f5b03f4d02768cf8d |
| ENCFF928WDI.tabix.bed.gz.tbi | 282.58 KB | 97fd4d1d1c82118a482c61e10c5b5f45 |
| ENCFF928WDI.tabix.bed.gz.tbi.dvc | 110.0 B | 4bf927f9349fcb4c76e4485224ff7694 |
| genomic_resource.yaml | 2.82 KB | 77f3eb08b33483ce1e32716d72909580 |
| genomic_resource_original.yaml | 2.66 KB | a94818cde91412b9cfb01f752f7fef6c |
| statistics/ |