Histone_ChIP-seq_ENCSR000AMT
| Id: | Histone_ChIP-seq/ENCSR000AMT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AMT [biosamplesummary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K36me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (45 years) female child (11 years) output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN169GIE|/analyses/ENCAN169GIE/} has in progress subobject document {d3408fc3-a914-40c3-aa65-df34ca0c3488|/documents/d3408fc3-a914-40c3-aa65-df34ca0c3488/} audit_internal_action: Released analysis {ENCAN169GIE|/analyses/ENCAN169GIE/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF569UUF|/files/ENCFF569UUF/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 7283646 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF757MMH|/files/ENCFF757MMH/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline has 13570447 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K36me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
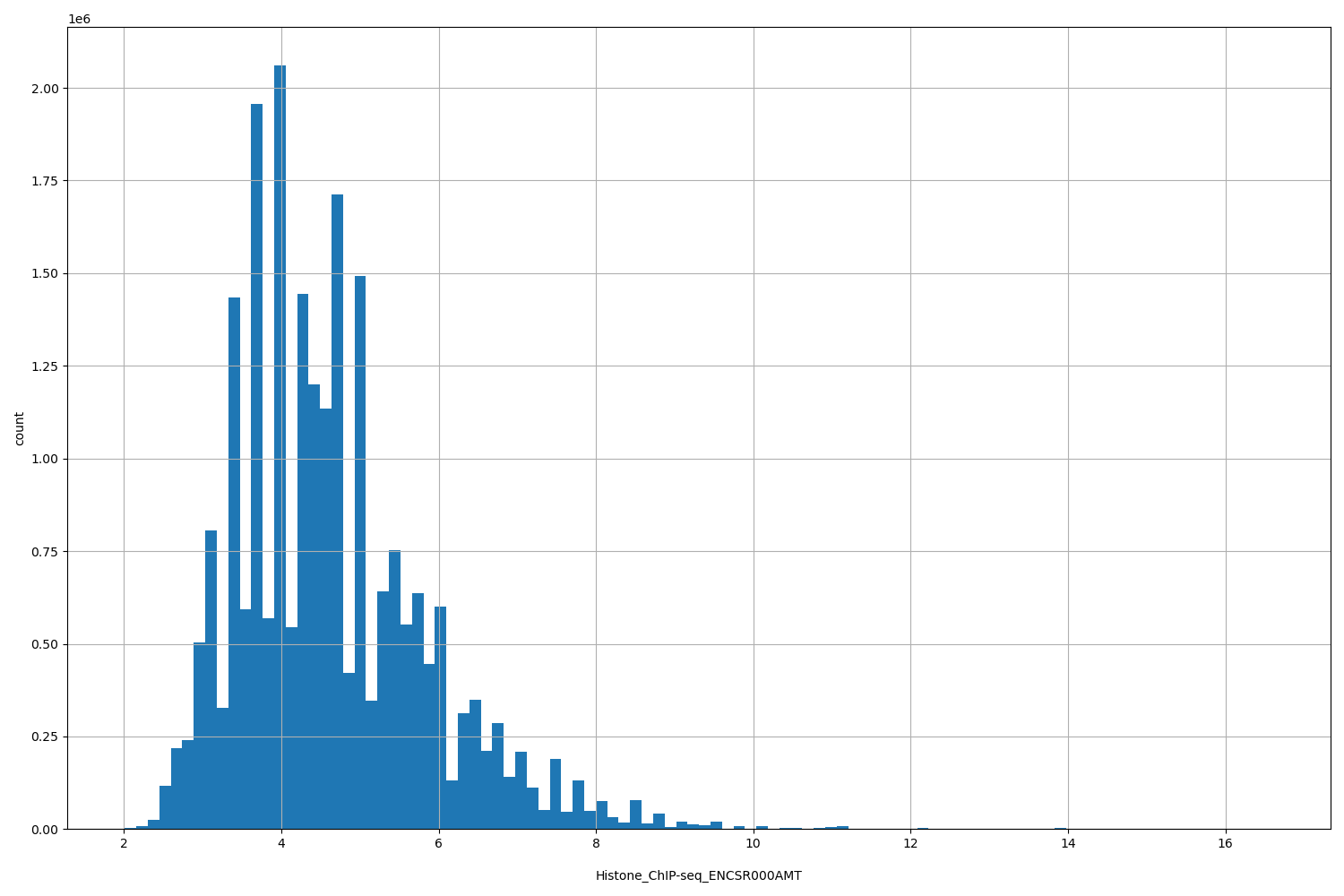
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AMT | float |
Histone_ChIP-seq_ENCSR000AMT |
Histone_ChIP-seq ENCSR000AMT [biosample_summary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K36me3"]
|
 |
[2.01, 16.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF722PLP.bed.gz | 1.28 MB | fb63f2da721533ae6e82988ca4331419 |
| ENCFF722PLP.bed.gz.dvc | 101.0 B | cd75776431455d5aa3e2036eaf7534bd |
| ENCFF722PLP.tabix.bed.gz | 704.77 KB | 43a1e43422627aa31201ecb40cc246bc |
| ENCFF722PLP.tabix.bed.gz.dvc | 106.0 B | abe6af8d3f1b6495d5fd7a91137053fc |
| ENCFF722PLP.tabix.bed.gz.tbi | 151.95 KB | 218bd305ef8b095db8789b288c010ecf |
| ENCFF722PLP.tabix.bed.gz.tbi.dvc | 110.0 B | d14e0f49c4c009fc62552f1f07f0f612 |
| genomic_resource.yaml | 2.84 KB | a12d6d9624ce4f44012417941f5b00fc |
| genomic_resource_original.yaml | 2.67 KB | 40ad8e94b0847556335ded6aa85970c4 |
| statistics/ |