Histone_ChIP-seq_ENCSR000AMR
| Id: | Histone_ChIP-seq/ENCSR000AMR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AMR [biosamplesummary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K27ac"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: male adult (45 years) female child (11 years) output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN227KII|/analyses/ENCAN227KII/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF377BNX|/files/ENCFF377BNX/} processed by ChIP-seq ENCODE3 hg19 pipeline has 7318543 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF066AQB|/files/ENCFF066AQB/} processed by ChIP-seq ENCODE3 hg19 pipeline has 6784781 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27ac-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
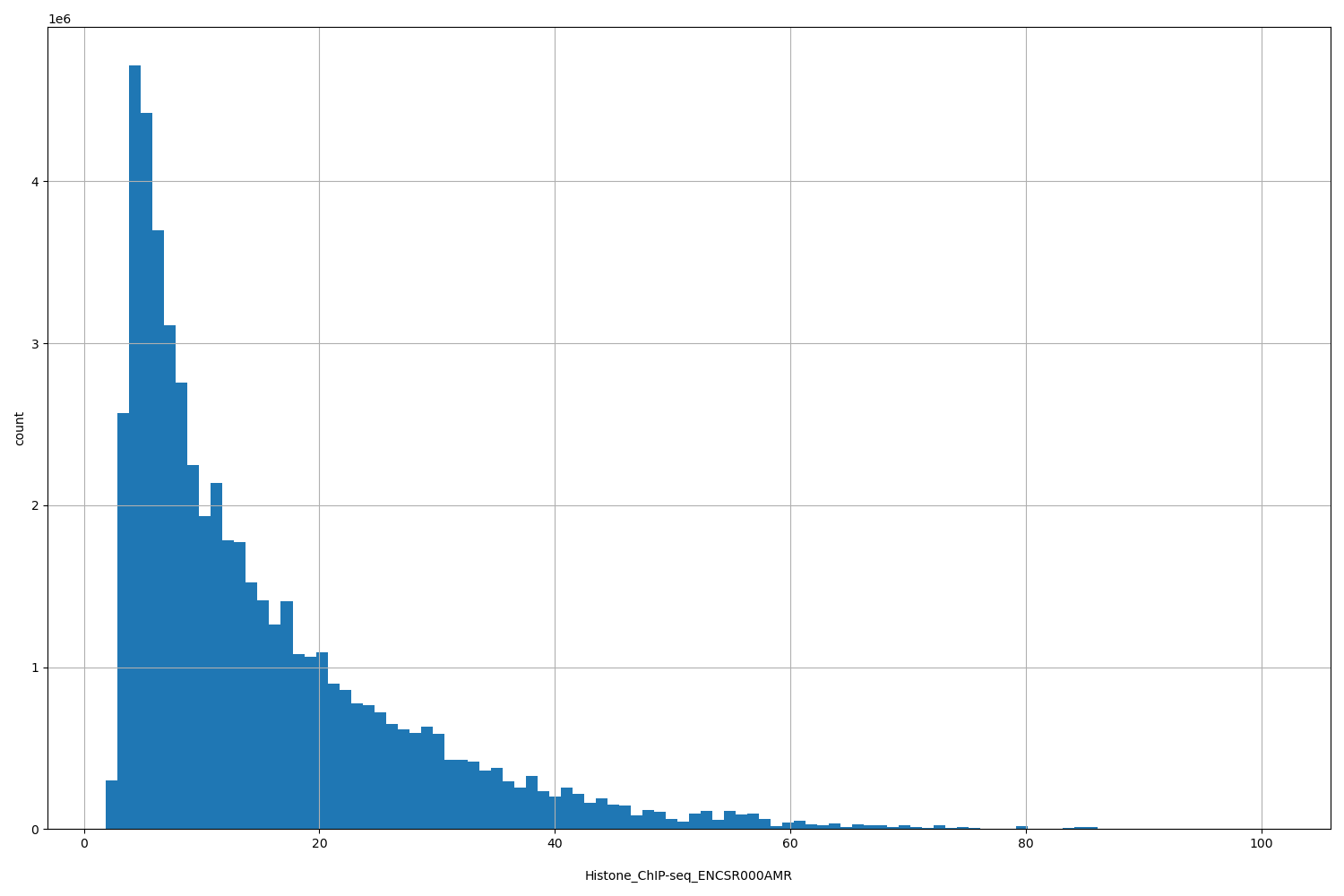
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AMR | float |
Histone_ChIP-seq_ENCSR000AMR |
Histone_ChIP-seq ENCSR000AMR [biosample_summary="Homo sapiens fibroblast of lung female child (11 years) and male adult (45 years)" and target="H3K27ac"]
|
 |
[1.87, 101] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF943QBE.bed.gz | 1.38 MB | a032af6a0f47925a834cab9624c28a4a |
| ENCFF943QBE.bed.gz.dvc | 101.0 B | a36d1650f54307372cd594cccf346848 |
| ENCFF943QBE.tabix.bed.gz | 664.07 KB | 590b0c3d3f26d267b8e6a684c42633d0 |
| ENCFF943QBE.tabix.bed.gz.dvc | 106.0 B | 43515e39a4b4a7d2b8e39e67b87b02a3 |
| ENCFF943QBE.tabix.bed.gz.tbi | 219.46 KB | 81aab7f1b9def353b966c157a8b5b483 |
| ENCFF943QBE.tabix.bed.gz.tbi.dvc | 110.0 B | bcf7c3edce0b825e0765e131098c067c |
| genomic_resource.yaml | 2.6 KB | 2d69da5f692ca1b64b0d875d0f017396 |
| genomic_resource_original.yaml | 2.43 KB | 18641c942e1a2fe6cff4033b21d5fc5a |
| statistics/ |