Histone_ChIP-seq_ENCSR000AMM
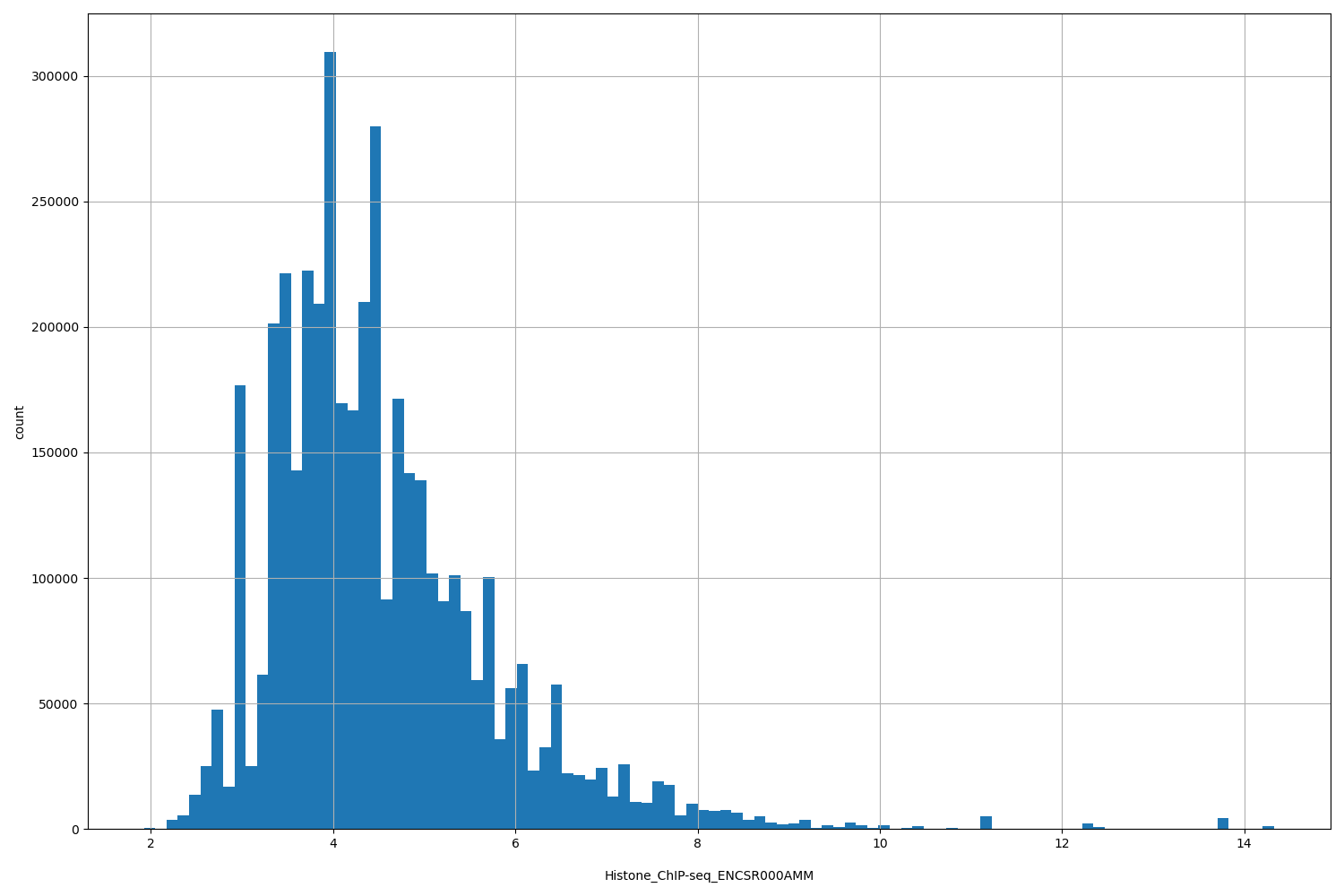
| Id: | Histone_ChIP-seq/ENCSR000AMM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AMM [biosamplesummary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="H4K20me1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 3 summary: female adult (50 years) output_type: replicated peaks audit_internal_action: Archived analysis {ENCAN601KTY|/analyses/ENCAN601KTY/} has in progress subobject quality standard {encode3-histone-chip|/quality-standards/encode3-histone-chip/} audit_not_compliant: Processed alignments file {ENCFF744MRF|/files/ENCFF744MRF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 8989881 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H4K20me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF120SIU|/files/ENCFF120SIU/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 11657189 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H4K20me1-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AMM | float |
Histone_ChIP-seq_ENCSR000AMM |
Histone_ChIP-seq ENCSR000AMM [biosample_summary="Homo sapiens mammary epithelial cell female adult (50 years)" and target="H4K20me1"]
|
 |
[1.92, 14.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF657XGG.bed.gz | 260.2 KB | 766b1f57f7c435f8259df0e40812992f |
| ENCFF657XGG.bed.gz.dvc | 100.0 B | 322027616be40e467f22cfcd5be0b4c1 |
| ENCFF657XGG.tabix.bed.gz | 131.91 KB | 157a443f95748e8b08cc66dc018ffaaa |
| ENCFF657XGG.tabix.bed.gz.dvc | 106.0 B | 585da11a740d351522a27df477e073cd |
| ENCFF657XGG.tabix.bed.gz.tbi | 71.17 KB | a523aac1dc077903002b7c6c36ce268e |
| ENCFF657XGG.tabix.bed.gz.tbi.dvc | 109.0 B | 45660d0ec0e967fa2457f6369fcf58e4 |
| genomic_resource.yaml | 2.49 KB | a0c7d2c1fc3f90b11238952f52f6ef09 |
| genomic_resource_original.yaml | 2.35 KB | b547822d78a42417dce19fad4257f6b2 |
| statistics/ |