Histone_ChIP-seq_ENCSR000AKT
| Id: | Histone_ChIP-seq/ENCSR000AKT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AKT [biosamplesummary="Homo sapiens K562" and target="H3K4me2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN608CZX|/analyses/ENCAN608CZX/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN608CZX|/analyses/ENCAN608CZX/} has in progress subobject document {2ded7c03-12ac-4d25-84e6-92286d54c460|/documents/2ded7c03-12ac-4d25-84e6-92286d54c460/} audit_not_compliant: Processed alignments file {ENCFF446FUS|/files/ENCFF446FUS/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9297178 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF583GSH|/files/ENCFF583GSH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6882054 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K4me2-human and investigated as a narrow histone mark is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
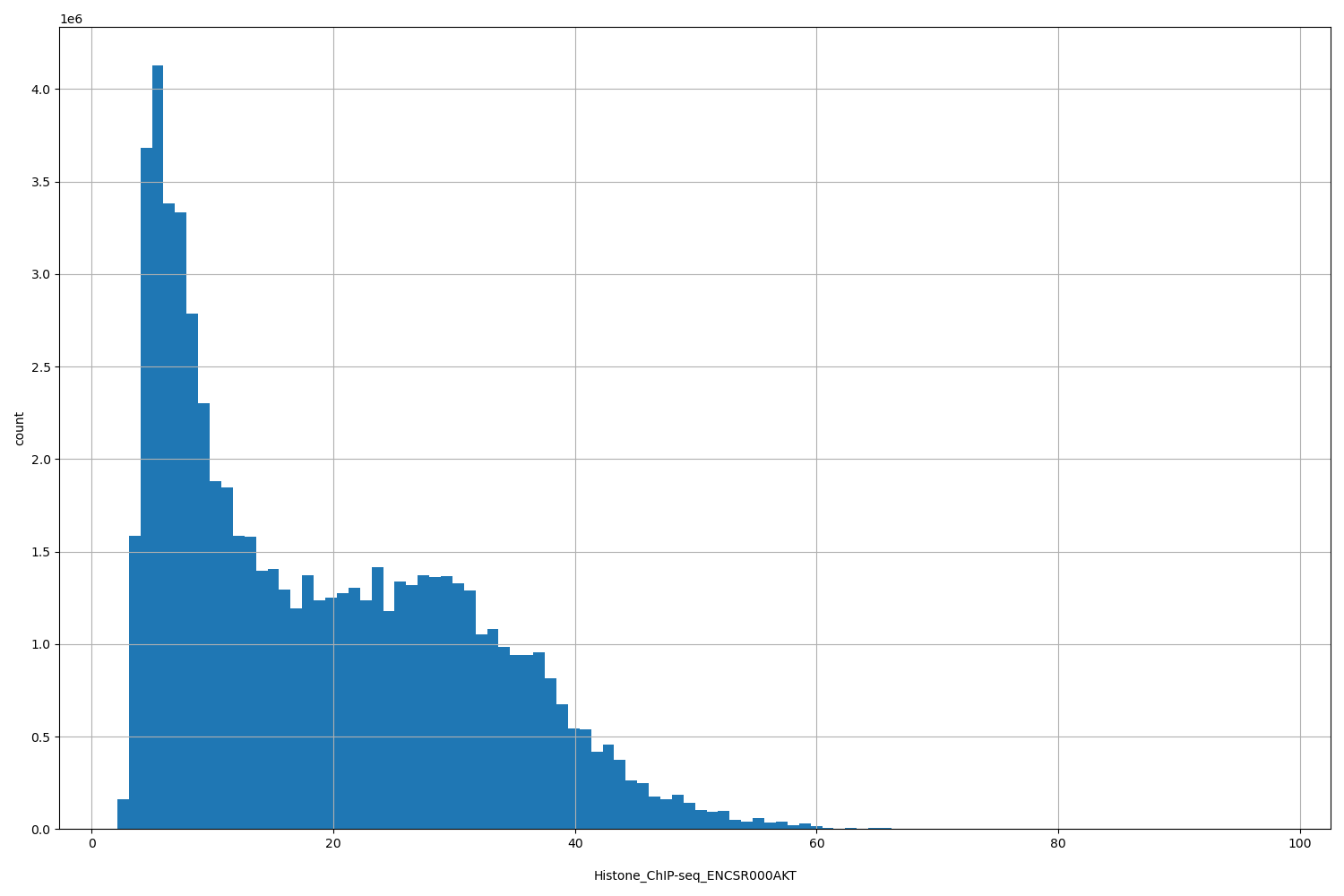
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AKT | float |
Histone_ChIP-seq_ENCSR000AKT |
Histone_ChIP-seq ENCSR000AKT [biosample_summary="Homo sapiens K562" and target="H3K4me2"]
|
 |
[2.13, 97.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF749KLQ.bed.gz | 1.58 MB | 55c1093b8d98bef1dc7f788fbb44eb7c |
| ENCFF749KLQ.bed.gz.dvc | 101.0 B | 58ce22b22df76c587b1f1db0acce8a49 |
| ENCFF749KLQ.tabix.bed.gz | 704.6 KB | 392c40c57a3ef07d1073d9be4f1b8db7 |
| ENCFF749KLQ.tabix.bed.gz.dvc | 106.0 B | 96149a6c28e5d89b6ff93b711a2dd03f |
| ENCFF749KLQ.tabix.bed.gz.tbi | 237.84 KB | 65d736a548f04ca3beb0737e294abee9 |
| ENCFF749KLQ.tabix.bed.gz.tbi.dvc | 110.0 B | 5cb5a08cef03acc5d56d05cf17217edf |
| genomic_resource.yaml | 2.6 KB | d25179e1c451457b6de2e064b80fa75b |
| genomic_resource_original.yaml | 2.5 KB | 04f7a2a6560736e5c91cf1ac04e8b038 |
| statistics/ |