Histone_ChIP-seq_ENCSR000AKD
| Id: | Histone_ChIP-seq/ENCSR000AKD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
HistoneChIP-seq ENCSR000AKD [biosamplesummary="Homo sapiens GM12878" and target="H3K27me3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: pseudoreplicated peaks audit_internal_action: Released analysis {ENCAN726MRC|/analyses/ENCAN726MRC/} has in progress subobject quality standard {encode4-histone-chip|/quality-standards/encode4-histone-chip/} audit_internal_action: Released analysis {ENCAN726MRC|/analyses/ENCAN726MRC/} has in progress subobject document {534d60ac-209b-4e4b-9ec6-e5205e4d1a84|/documents/534d60ac-209b-4e4b-9ec6-e5205e4d1a84/} audit_not_compliant: Processed alignments file {ENCFF633BHN|/files/ENCFF633BHN/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 10101892 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) audit_not_compliant: Processed alignments file {ENCFF565DCK|/files/ENCFF565DCK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6346970 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting H3K27me3-human and investigated as a broad histone mark is 20 million usable fragments. The recommended value is > 45 million, but > 35 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/}) |
| Labels: |
|
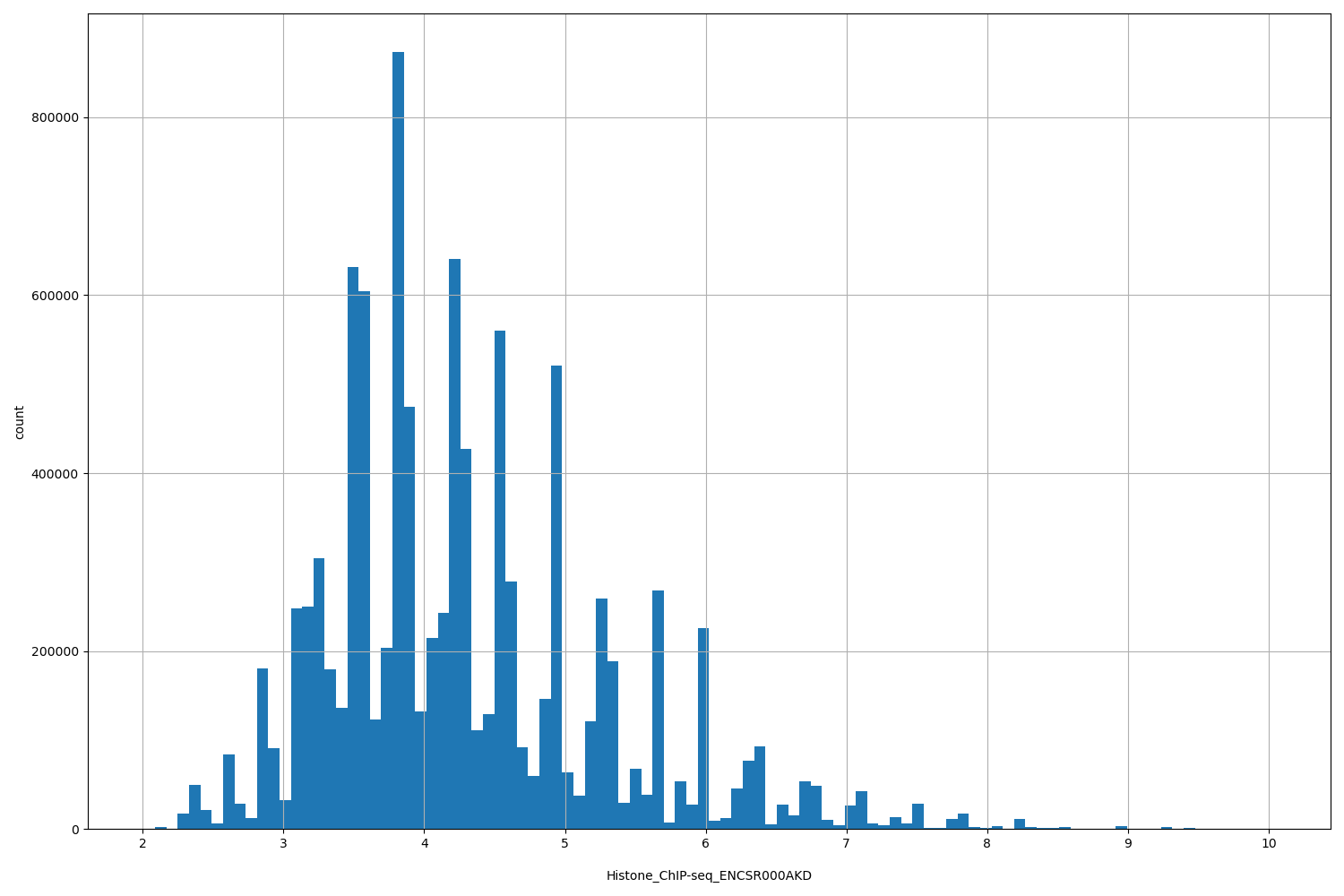
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| Histone_ChIP-seq_ENCSR000AKD | float |
Histone_ChIP-seq_ENCSR000AKD |
Histone_ChIP-seq ENCSR000AKD [biosample_summary="Homo sapiens GM12878" and target="H3K27me3"]
|
 |
[2.01, 10] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF291DHI.bed.gz | 420.41 KB | 7de99dd8ded03be57595a07474f34ebb |
| ENCFF291DHI.bed.gz.dvc | 100.0 B | f633beaa0d9395d77e352673aa0d609a |
| ENCFF291DHI.tabix.bed.gz | 235.57 KB | 0c7da97d31f37137a6a404757f31c299 |
| ENCFF291DHI.tabix.bed.gz.dvc | 106.0 B | 2799328345cf5897f2638d1a2c8a4e81 |
| ENCFF291DHI.tabix.bed.gz.tbi | 123.22 KB | 2381d2778363472cce58b3aa1a7644f9 |
| ENCFF291DHI.tabix.bed.gz.tbi.dvc | 110.0 B | 0e6ae35c383685498b0008eb90e51587 |
| genomic_resource.yaml | 2.62 KB | 73f1f04869d340662bf66ed6b67b141e |
| genomic_resource_original.yaml | 2.51 KB | 3fb0761ce51ca26f7cf1dd56957808fa |
| statistics/ |