DNase-seq_ENCSR970DQR
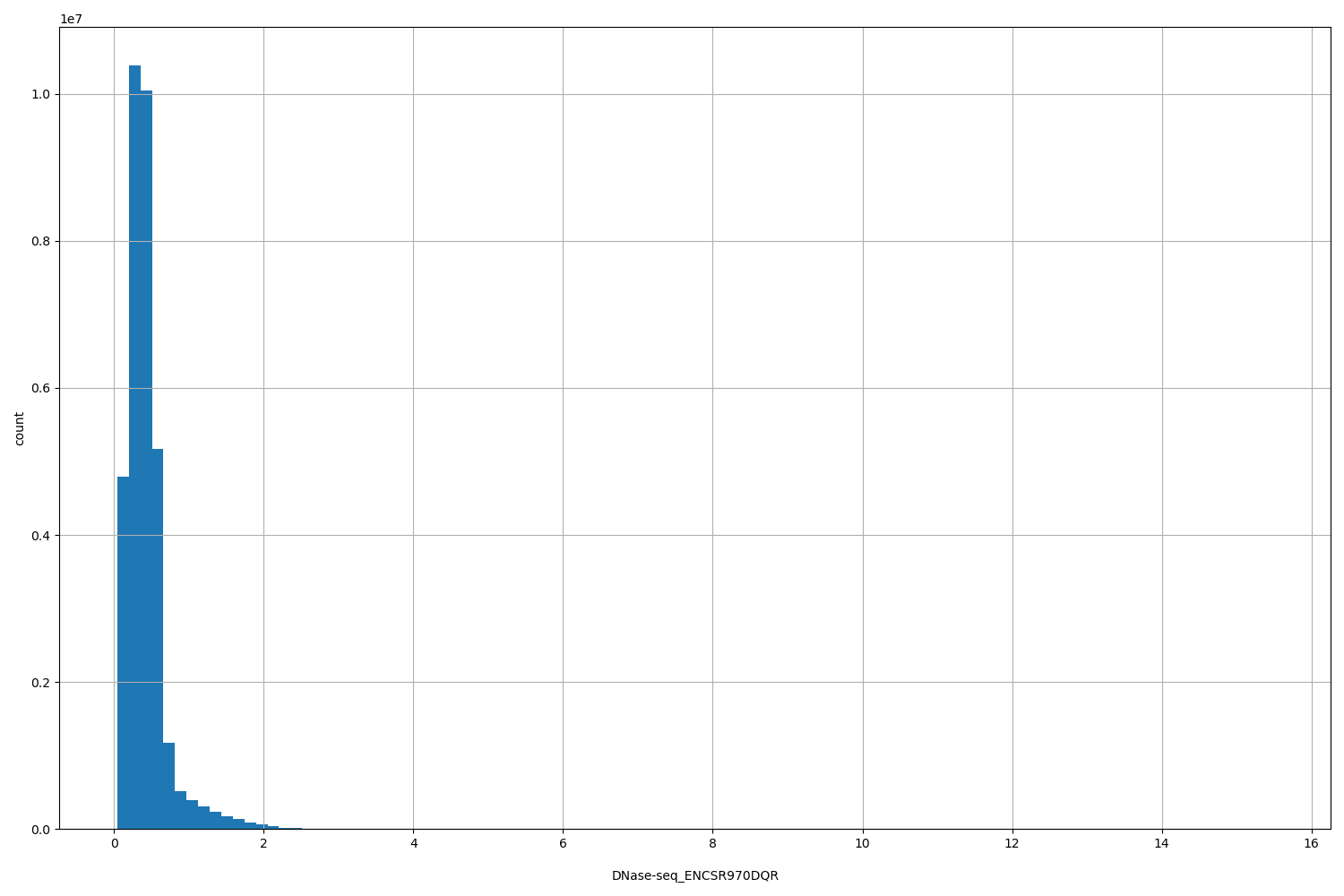
| Id: | DNase-seq/ENCSR970DQR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR970DQR [biosample_summary="Homo sapiens chondrocyte"] |
| Description: |
status: released biological_replicates: Rep 2 summary: female embryo (5 days) output_type: peaks audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF098QWQ|/files/ENCFF098QWQ/} processed by DNase-seq ENCODE4 v3.0.0 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.32. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF099TKH|/files/ENCFF099TKH/} processed by DNase-seq ENCODE4 v3.0.0 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.34. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR970DQR | float |
DNase-seq_ENCSR970DQR |
DNase-seq ENCSR970DQR [biosample_summary="Homo sapiens chondrocyte"]
|
 |
[0.0444, 15.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF429PXE.bed.gz | 1.45 MB | b5c45f5105517b46f976ace29b123548 |
| ENCFF429PXE.bed.gz.dvc | 101.0 B | 99c912adcb217515c035fa5e1148b82d |
| ENCFF429PXE.tabix.bed.gz | 1.4 MB | ab101c4694828bd06074c96b6fddaef0 |
| ENCFF429PXE.tabix.bed.gz.dvc | 107.0 B | eb890534671ae52931e40bb3301adf9b |
| ENCFF429PXE.tabix.bed.gz.tbi | 218.38 KB | 543855e743764a6d850f1a66afb2958d |
| ENCFF429PXE.tabix.bed.gz.tbi.dvc | 110.0 B | b2cf4b294dc753ab31b2dac85c80baca |
| genomic_resource.yaml | 2.77 KB | 8c3a52a1f32b462f7b1a39265a6dfe03 |
| genomic_resource_original.yaml | 2.7 KB | 0618f77e4264e78d1790c787748c8ef8 |
| statistics/ |