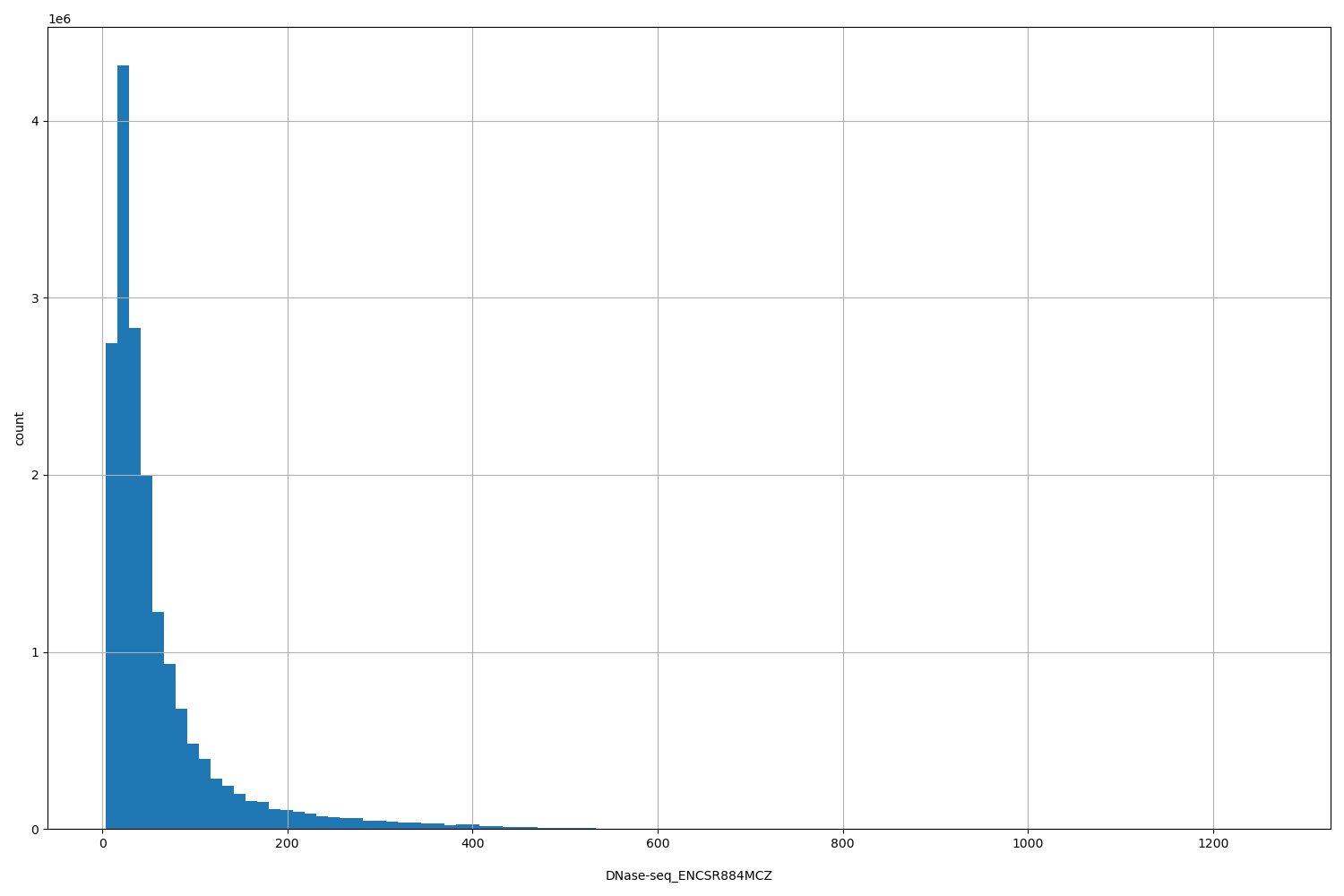
DNase-seq_ENCSR884MCZ
| Id: | DNase-seq/ENCSR884MCZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR884MCZ [biosample_summary="Homo sapiens right kidney tissue female embryo (98 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (98 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN654JBV|/analyses/ENCAN654JBV/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF376JOV|/files/ENCFF376JOV/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 42365277 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF614MII|/files/ENCFF614MII/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 42097469 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR884MCZ | float |
DNase-seq_ENCSR884MCZ |
DNase-seq ENCSR884MCZ [biosample_summary="Homo sapiens right kidney tissue female embryo (98 days)"]
|
 |
[4, 1.26e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF717QSC.bed.gz | 1.12 MB | dd5ae833cb6671a6afcf5149945d08f9 |
| ENCFF717QSC.bed.gz.dvc | 101.0 B | 82b17ba89486bfef0afa973122f7c283 |
| ENCFF717QSC.tabix.bed.gz | 981.47 KB | 83f9a81ad6f813aa67825fe5220f99b8 |
| ENCFF717QSC.tabix.bed.gz.dvc | 107.0 B | 360026aaa2a2a6386c0e0f880042db43 |
| ENCFF717QSC.tabix.bed.gz.tbi | 528.36 KB | d1ba6c1d83ce44a38347693a66f3c566 |
| ENCFF717QSC.tabix.bed.gz.tbi.dvc | 110.0 B | 637dea629c2ab210f113606d138c96b6 |
| genomic_resource.yaml | 2.6 KB | be8e7f05bcaab7c1856d92d04ba2517a |
| genomic_resource_original.yaml | 2.5 KB | 8150c64706b6fea1b2bd8b9e4d40a8e7 |
| statistics/ |