DNase-seq_ENCSR814KRX
| Id: | DNase-seq/ENCSR814KRX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR814KRX [biosample_summary="Homo sapiens HT-29"] |
| Description: |
status: released biological_replicates: Rep 2 summary: output_type: peaks audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF698NHT|/files/ENCFF698NHT/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.38. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
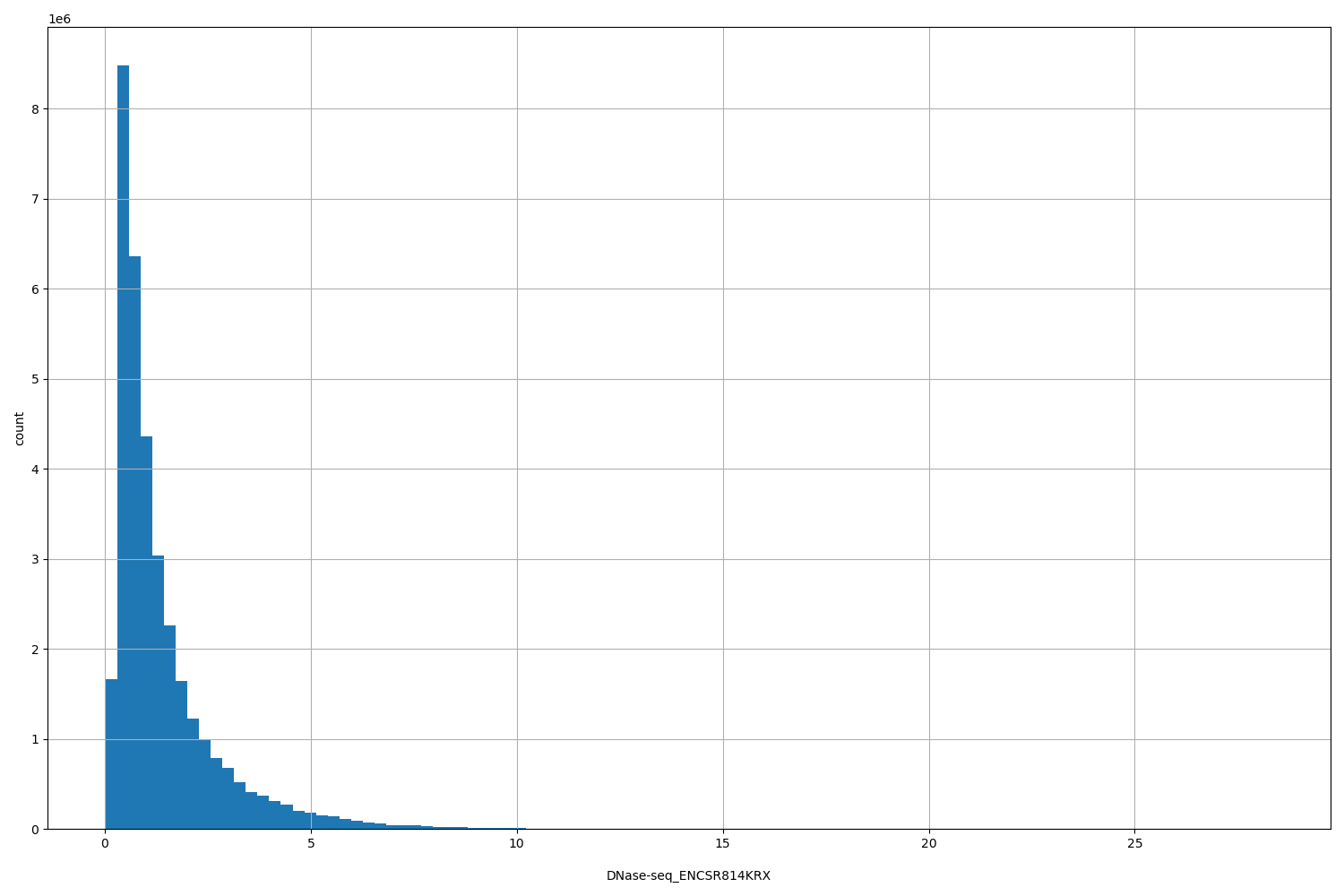
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR814KRX | float |
DNase-seq_ENCSR814KRX |
DNase-seq ENCSR814KRX [biosample_summary="Homo sapiens HT-29"]
|
 |
[0.0257, 28.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF094XEJ.bed.gz | 1.67 MB | 5b771762f1fc5db71e76d31311f2fc38 |
| ENCFF094XEJ.bed.gz.dvc | 101.0 B | 2eb2754874feb6be8342f28d350ac7ee |
| ENCFF094XEJ.tabix.bed.gz | 1.56 MB | 87b23ed1a1fee6f531d453a3c8ed6a72 |
| ENCFF094XEJ.tabix.bed.gz.dvc | 107.0 B | 4115830677e039e371ea9bdc11a06615 |
| ENCFF094XEJ.tabix.bed.gz.tbi | 524.29 KB | abee00e7734001f673918b6679f4da93 |
| ENCFF094XEJ.tabix.bed.gz.tbi.dvc | 110.0 B | 8c4fe559f8ca98c62021b9094b1b4265 |
| genomic_resource.yaml | 1.91 KB | 1bd884cbea8f4f76535284602a1337ca |
| genomic_resource_original.yaml | 1.84 KB | b072bc17958fa0dcc02d61636fb9a3f6 |
| statistics/ |