DNase-seq_ENCSR765BSU
| Id: | DNase-seq/ENCSR765BSU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR765BSU [biosample_summary="Homo sapiens right kidney tissue female embryo (117 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (117 days) output_type: peaks audit_warning: Alignment file {ENCFF907UIS|/files/ENCFF907UIS/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 34748745 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF907UIS|/files/ENCFF907UIS/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.33. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
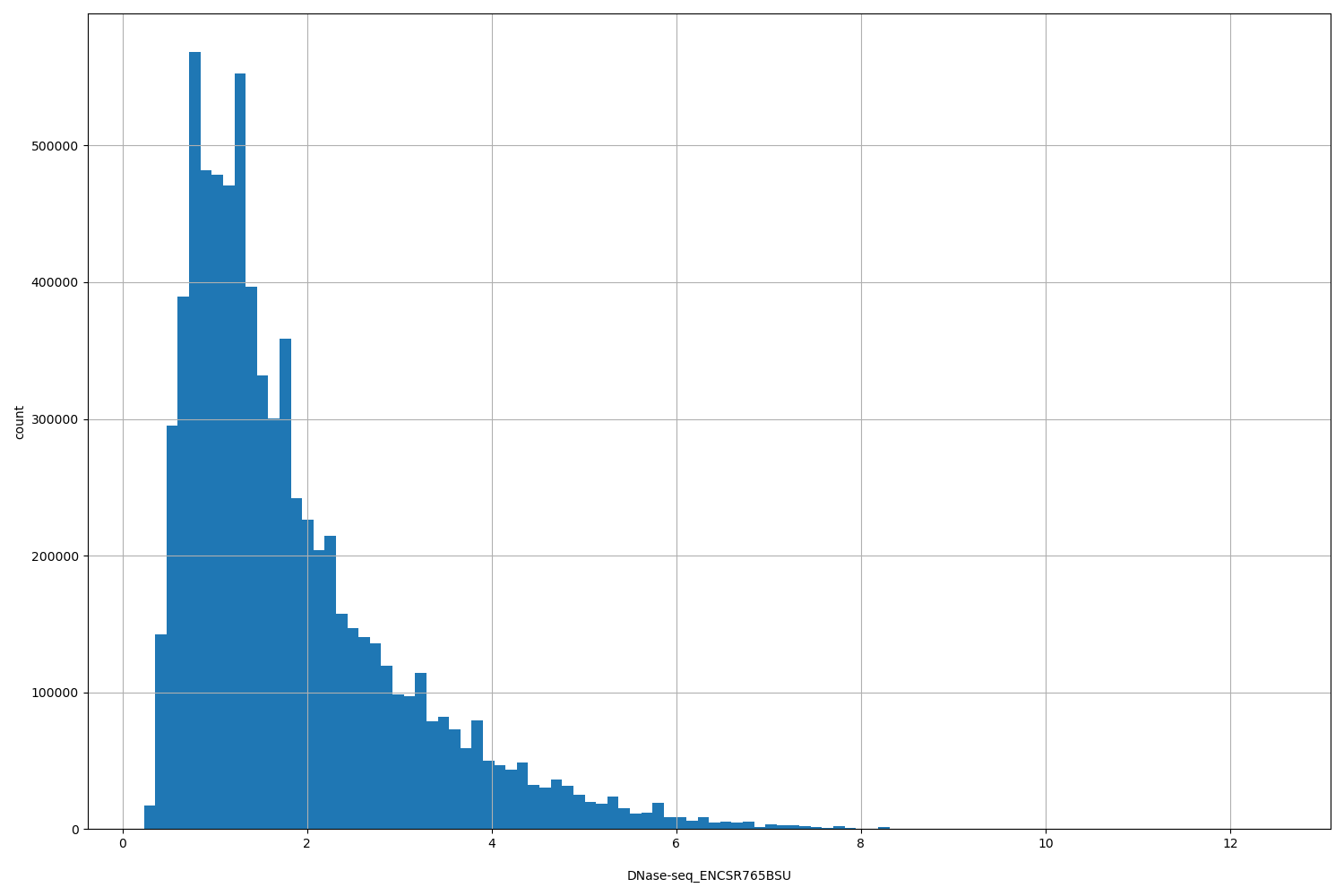
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR765BSU | float |
DNase-seq_ENCSR765BSU |
DNase-seq ENCSR765BSU [biosample_summary="Homo sapiens right kidney tissue female embryo (117 days)"]
|
 |
[0.231, 12.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF939AKZ.bed.gz | 392.93 KB | 9ef1e4bea2eb31c0669edf340011c767 |
| ENCFF939AKZ.bed.gz.dvc | 100.0 B | 1785e11734edf30fa3f5358aa85d128d |
| ENCFF939AKZ.tabix.bed.gz | 356.97 KB | 709b41d9c749a8cc981dd2a5f1a310da |
| ENCFF939AKZ.tabix.bed.gz.dvc | 106.0 B | 83b88f3989533281d19c7bf171dfd9c9 |
| ENCFF939AKZ.tabix.bed.gz.tbi | 207.76 KB | 3c1c2b2a3e7af96c7438aa1da9b72111 |
| ENCFF939AKZ.tabix.bed.gz.tbi.dvc | 110.0 B | 1537964cf6b5ec1c060f60e23a172844 |
| genomic_resource.yaml | 2.67 KB | 4400c29cdb6df96b49af6021193036e5 |
| genomic_resource_original.yaml | 2.57 KB | 41922446b4281bb5c9dd669c34bfd692 |
| statistics/ |