DNase-seq_ENCSR729DRB
| Id: | DNase-seq/ENCSR729DRB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR729DRB [biosample_summary="Homo sapiens testis tissue male embryo"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo output_type: peaks audit_internal_action: Archived analysis {ENCAN356ROA|/analyses/ENCAN356ROA/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF089BVK|/files/ENCFF089BVK/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 43243655 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF188DIT|/files/ENCFF188DIT/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 42923516 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
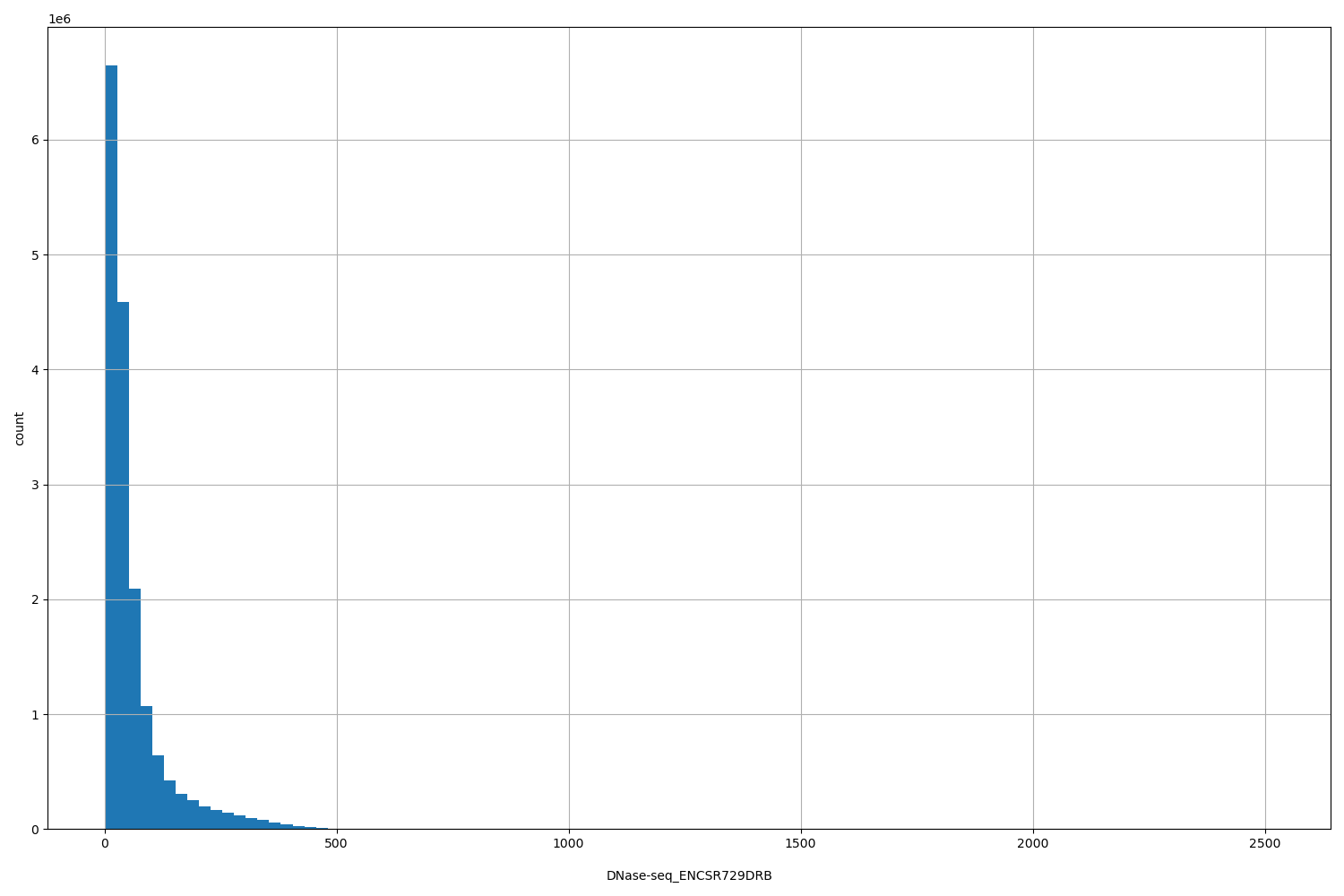
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR729DRB | float |
DNase-seq_ENCSR729DRB |
DNase-seq ENCSR729DRB [biosample_summary="Homo sapiens testis tissue male embryo"]
|
 |
[3, 2.52e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF618EIJ.bed.gz | 1.07 MB | 750f48bda5fc29c49152ddca38932dd1 |
| ENCFF618EIJ.bed.gz.dvc | 101.0 B | a7dedae9e78da23e85ee26a321daa250 |
| ENCFF618EIJ.tabix.bed.gz | 939.02 KB | 61028a8463751f613b7a0541aa1fd033 |
| ENCFF618EIJ.tabix.bed.gz.dvc | 106.0 B | 9d62ed7522874e5fa07146de6045e878 |
| ENCFF618EIJ.tabix.bed.gz.tbi | 502.64 KB | 1012637a7069d147dca60d63a926fe88 |
| ENCFF618EIJ.tabix.bed.gz.tbi.dvc | 110.0 B | 29993590f846697373f7074ba7e0de4a |
| genomic_resource.yaml | 2.52 KB | a866eb0948b05fde90751c742dd4f237 |
| genomic_resource_original.yaml | 2.43 KB | 98d4fd3cca261dffb8f9f77635599155 |
| statistics/ |