DNase-seq_ENCSR693UHT
| Id: | DNase-seq/ENCSR693UHT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR693UHT [biosample_summary="Homo sapiens omental fat pad tissue female adult (51 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (51 years) output_type: peaks audit_internal_action: Archived analysis {ENCAN615IWQ|/analyses/ENCAN615IWQ/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF344IWY|/files/ENCFF344IWY/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL202DNS|/pipelines/ENCPL202DNS/} ) for GRCh38 assembly has 39095848 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
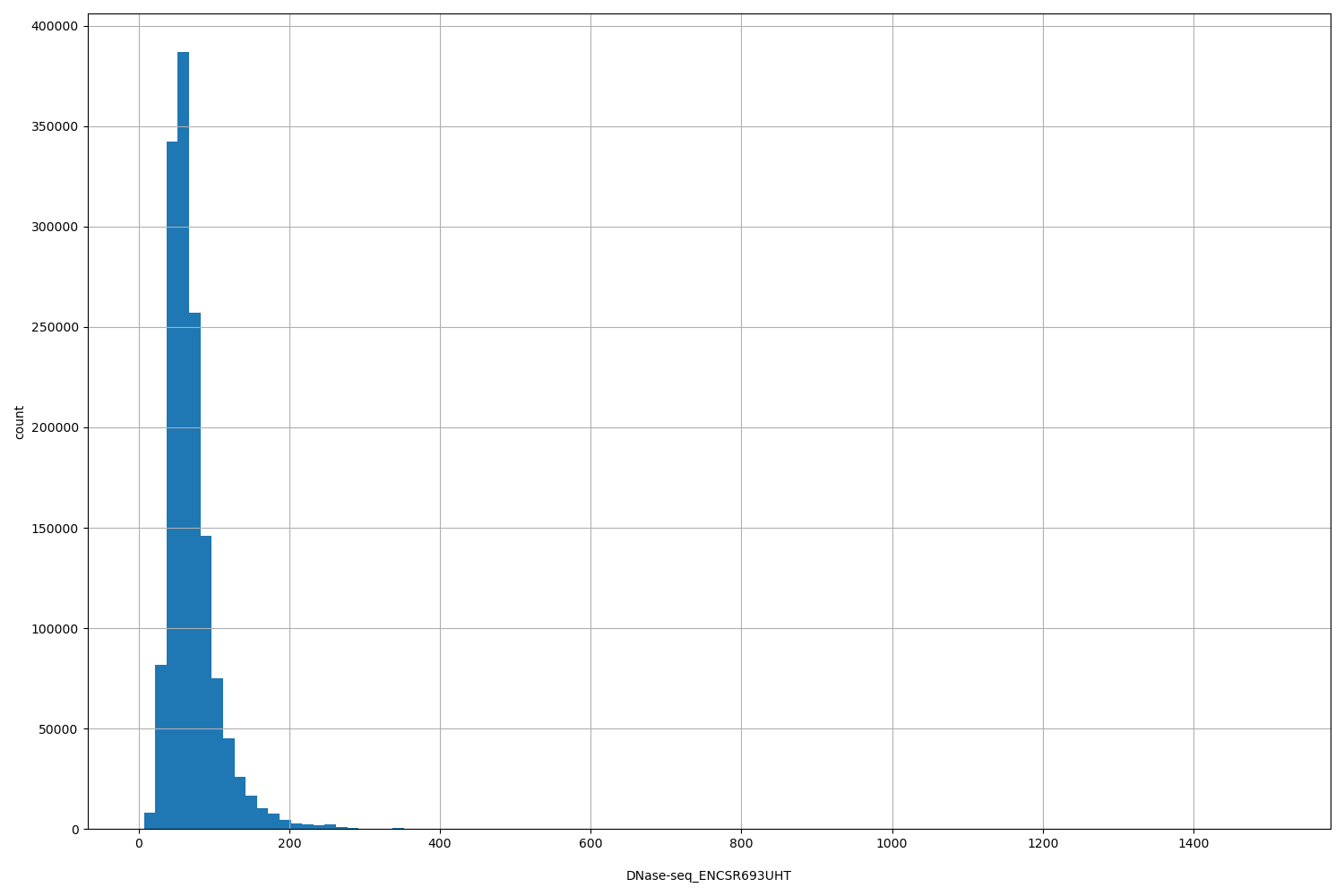
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR693UHT | float |
DNase-seq_ENCSR693UHT |
DNase-seq ENCSR693UHT [biosample_summary="Homo sapiens omental fat pad tissue female adult (51 years)"]
|
 |
[7, 1.51e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF188LZJ.bed.gz | 99.44 KB | 3635b5d6f504d70a271761731d25adbf |
| ENCFF188LZJ.bed.gz.dvc | 100.0 B | 1d004ed839821ca6ded9c8fb99e8a13f |
| ENCFF188LZJ.tabix.bed.gz | 83.05 KB | 7568fcb266d4b7ebc36a10b3f47df189 |
| ENCFF188LZJ.tabix.bed.gz.dvc | 105.0 B | 223f3a9d4c424b3d244767668172122c |
| ENCFF188LZJ.tabix.bed.gz.tbi | 77.79 KB | 29792f750dc7afc1e8770c1c48732652 |
| ENCFF188LZJ.tabix.bed.gz.tbi.dvc | 109.0 B | 88776a7165b58316a80f7ce2252930fc |
| genomic_resource.yaml | 2.01 KB | 648fd1381ada3d65ba70985de89e7ac1 |
| genomic_resource_original.yaml | 1.9 KB | 2c89c7d8c6346d9e76eb728cf652309c |
| statistics/ |