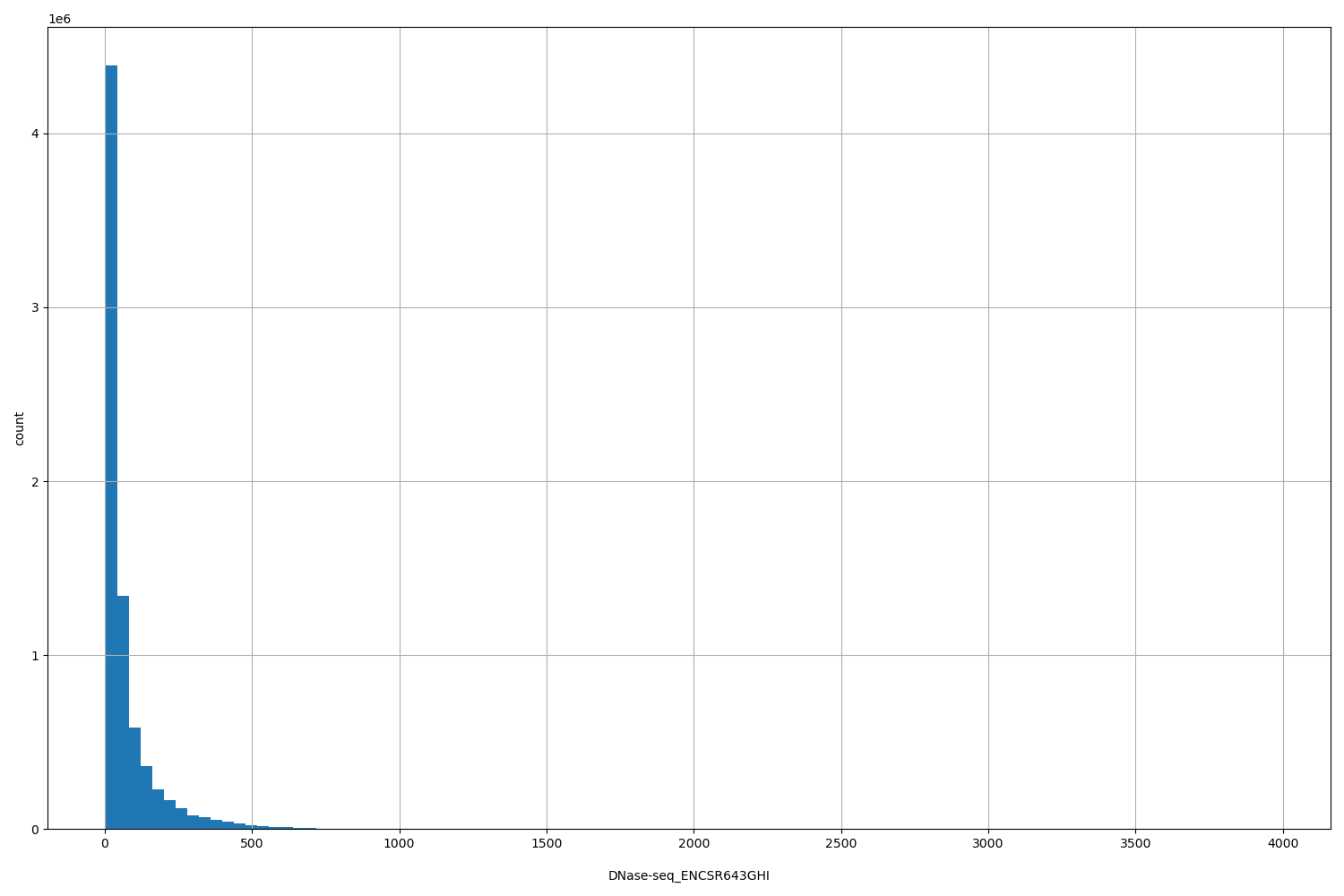
DNase-seq_ENCSR643GHI
| Id: | DNase-seq/ENCSR643GHI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR643GHI [biosample_summary="Homo sapiens CD4-positive, alpha-beta T cell female adult (33 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (33 years) output_type: peaks audit_internal_action: Archived analysis {ENCAN401SUC|/analyses/ENCAN401SUC/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF787DAE|/files/ENCFF787DAE/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 32491105 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF263RQE|/files/ENCFF263RQE/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 32252901 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR643GHI | float |
DNase-seq_ENCSR643GHI |
DNase-seq ENCSR643GHI [biosample_summary="Homo sapiens CD4-positive, alpha-beta T cell female adult (33 years)"]
|
 |
[4, 3.96e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF412EJZ.bed.gz | 493.14 KB | a9a75258f1162ae474650f315296993d |
| ENCFF412EJZ.bed.gz.dvc | 100.0 B | 895344cfeb4777e3670a8734cd41fb0a |
| ENCFF412EJZ.tabix.bed.gz | 421.05 KB | 9e510b106ebaf978113ced017222c04a |
| ENCFF412EJZ.tabix.bed.gz.dvc | 106.0 B | bf62636d61bab7f90cf02360dc9bba73 |
| ENCFF412EJZ.tabix.bed.gz.tbi | 256.81 KB | b45f6adc0e8777794420deb74b183720 |
| ENCFF412EJZ.tabix.bed.gz.tbi.dvc | 110.0 B | 18a099261972e6112c88c4964d05812e |
| genomic_resource.yaml | 2.64 KB | 4733e683da663aab29fd23f94d483664 |
| genomic_resource_original.yaml | 2.53 KB | 535a6a0e90da5ea0771756798d070c8a |
| statistics/ |