DNase-seq_ENCSR600RAA
| Id: | DNase-seq/ENCSR600RAA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR600RAA [biosample_summary="Homo sapiens left kidney tissue male embryo (87 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (87 days) output_type: peaks audit_warning: Alignment file {ENCFF906FBZ|/files/ENCFF906FBZ/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 44872711 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
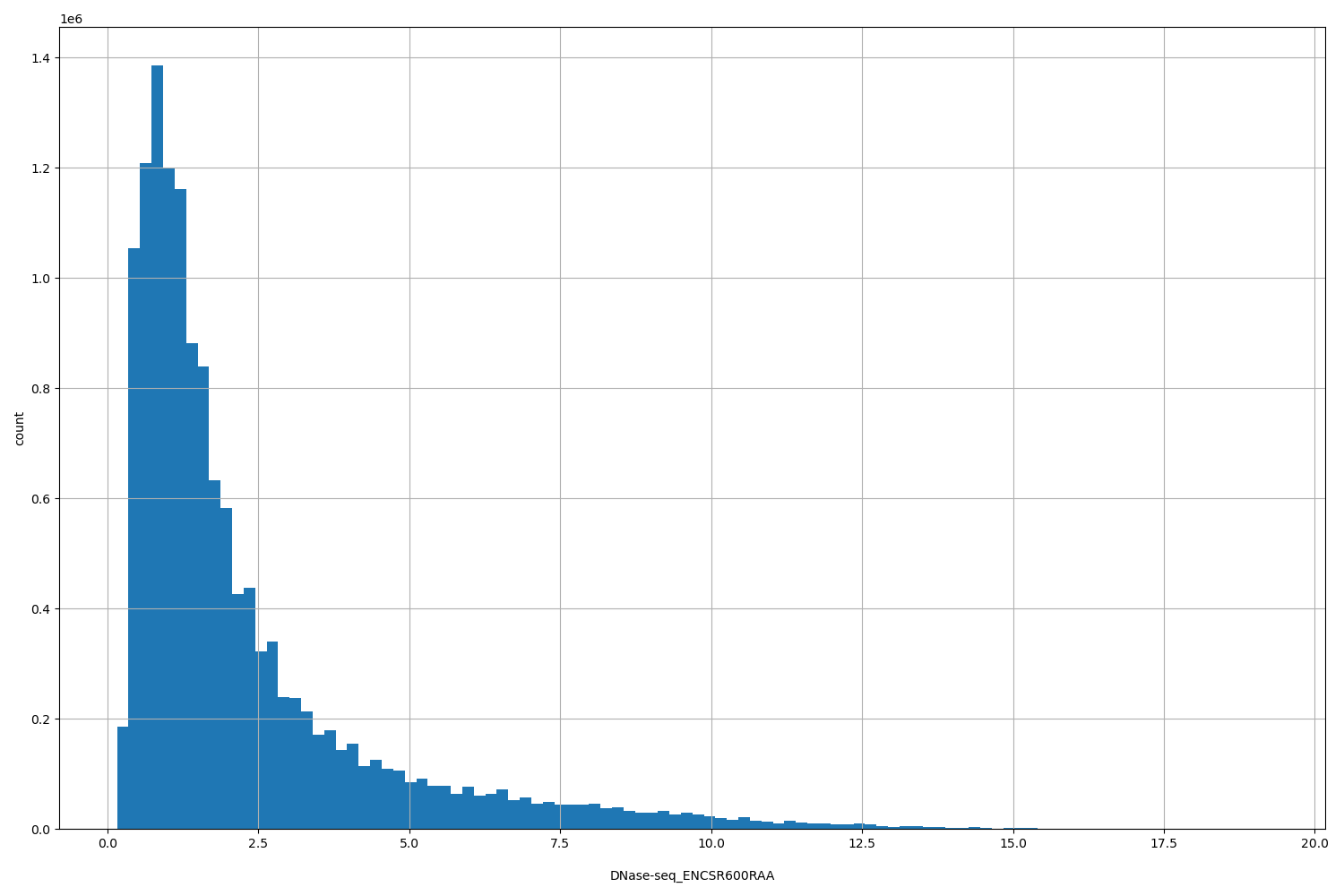
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR600RAA | float |
DNase-seq_ENCSR600RAA |
DNase-seq ENCSR600RAA [biosample_summary="Homo sapiens left kidney tissue male embryo (87 days)"]
|
 |
[0.156, 19.2] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF991MBB.bed.gz | 763.41 KB | aac53d1afa6b9f67c858bf4b6d4ea050 |
| ENCFF991MBB.bed.gz.dvc | 100.0 B | ce6e4f94c032b95490070dbed438f0a5 |
| ENCFF991MBB.tabix.bed.gz | 691.63 KB | 483c280002514a6f83341ceeaf119076 |
| ENCFF991MBB.tabix.bed.gz.dvc | 106.0 B | f1fd98995f4c14420e95d2a54df4d144 |
| ENCFF991MBB.tabix.bed.gz.tbi | 352.25 KB | c0412f861c9cafc67ed60c2e01bdf315 |
| ENCFF991MBB.tabix.bed.gz.tbi.dvc | 110.0 B | e1fcf80f444c78d577a50f7abc557413 |
| genomic_resource.yaml | 1.83 KB | 25a2ca6200f040d7ab00d745cff7bd5b |
| genomic_resource_original.yaml | 1.73 KB | 41b586c0eef47179f45dda081170fb71 |
| statistics/ |