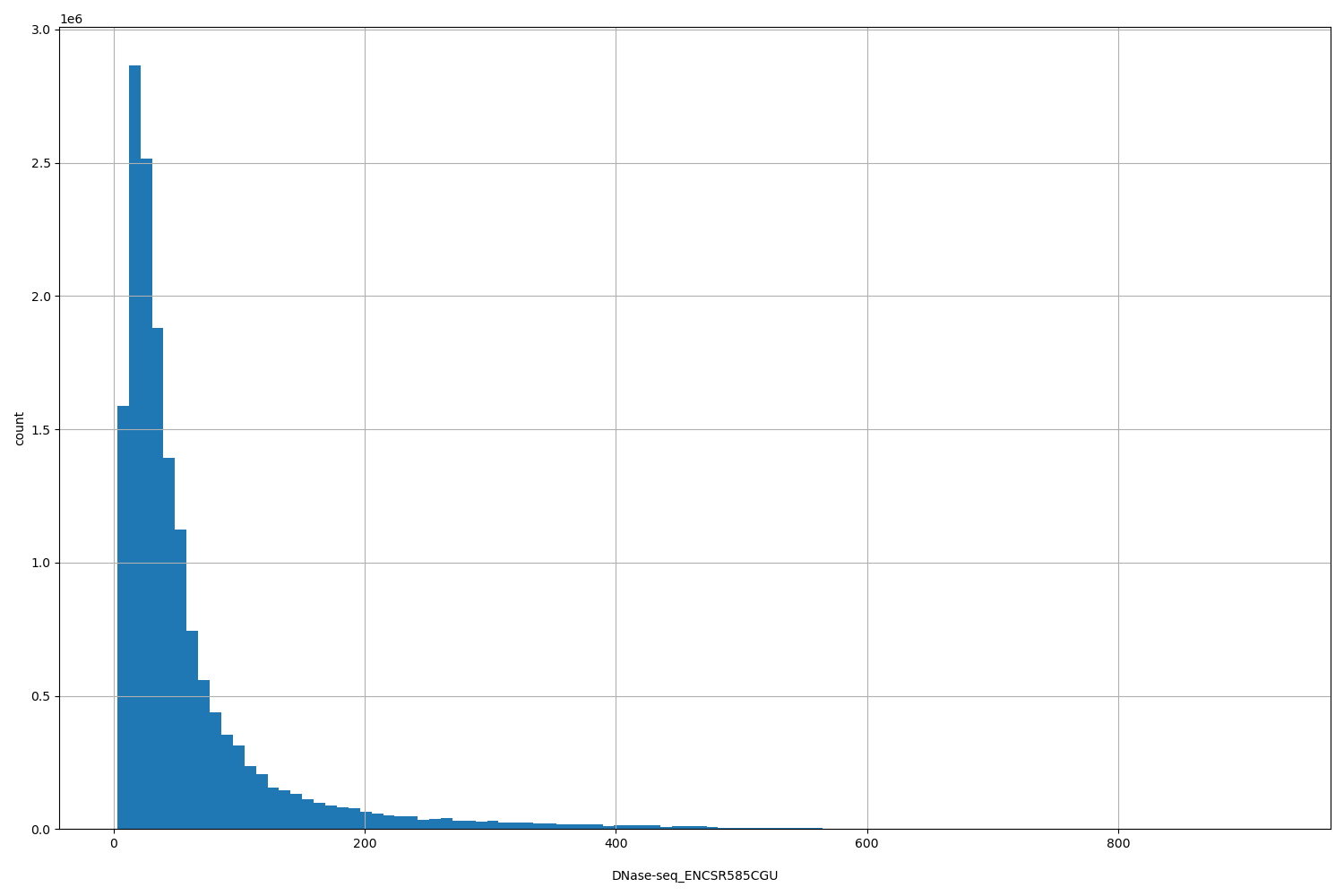
DNase-seq_ENCSR585CGU
| Id: | DNase-seq/ENCSR585CGU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR585CGU [biosample_summary="Homo sapiens muscle of back tissue female embryo (105 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (105 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN668SLL|/analyses/ENCAN668SLL/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF351FIS|/files/ENCFF351FIS/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 33553658 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF695PEK|/files/ENCFF695PEK/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 33342609 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR585CGU | float |
DNase-seq_ENCSR585CGU |
DNase-seq ENCSR585CGU [biosample_summary="Homo sapiens muscle of back tissue female embryo (105 days)"]
|
 |
[3, 923] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF897TZA.bed.gz | 1.01 MB | 953b478260b45d09798e4551cb141d36 |
| ENCFF897TZA.bed.gz.dvc | 101.0 B | 53747d36fc814a754a3d7f973edebdc5 |
| ENCFF897TZA.tabix.bed.gz | 886.21 KB | 730bcae1e917fe6ca39cd78ba659134f |
| ENCFF897TZA.tabix.bed.gz.dvc | 106.0 B | 2bebd379edbe8f154362cb12fbe6ac99 |
| ENCFF897TZA.tabix.bed.gz.tbi | 499.08 KB | 93c09f4c93dfec9ba23c67bdadd7d45b |
| ENCFF897TZA.tabix.bed.gz.tbi.dvc | 110.0 B | 43728457cbaf62451f47f13400e672be |
| genomic_resource.yaml | 2.61 KB | 8a6e266f504f578f94a37917c847b623 |
| genomic_resource_original.yaml | 2.5 KB | d11ad42c0e2c0c7c246d07c358edd65a |
| statistics/ |