DNase-seq_ENCSR566HLQ
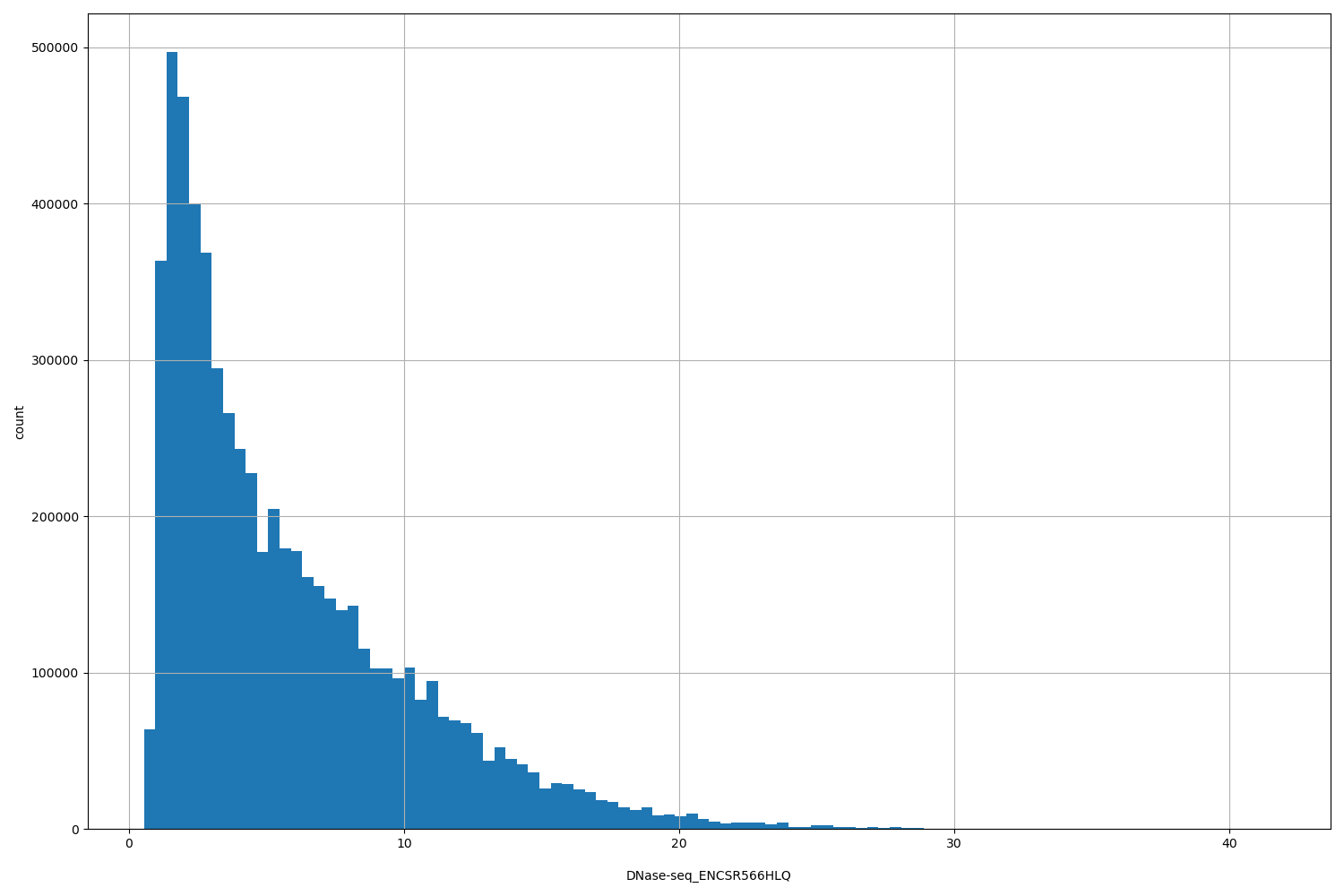
| Id: | DNase-seq/ENCSR566HLQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR566HLQ [biosample_summary="Homo sapiens stimulated activated CD4-positive, alpha-beta T cell female adult (33 years) treated with anti-CD3 and anti-CD28 coated beads for 24 hours, 100 ng/mL Interleukin-23 for 24 hours, 100 ng/mL Interleukin-1b for 24 hours"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (33 years) treated with anti-CD3 and anti-CD28 coated beads for 24 hours, 100 ng/mL Interleukin-23 for 24 hours, 100 ng/mL Interleukin-1b for 24 hours output_type: peaks audit_warning: Alignment file {ENCFF867LDP|/files/ENCFF867LDP/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 24146668 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR566HLQ | float |
DNase-seq_ENCSR566HLQ |
DNase-seq ENCSR566HLQ [biosample_summary="Homo sapiens stimulated activated CD4-positive, alpha-beta T cell female adult (33 years) treated with anti-CD3 and anti-CD28 coated beads for 24 hours, 100 ng/mL Interleukin-23 for 24 hours, 100 ng/mL Interleukin-1b for 24 hours"]
|
 |
[0.54, 41.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF792WBY.bed.gz | 361.09 KB | 8b77eb0bc4b002b8321ff2cd176ff28e |
| ENCFF792WBY.bed.gz.dvc | 100.0 B | 015a42f3ff317012a8ec15623eb28aad |
| ENCFF792WBY.tabix.bed.gz | 327.0 KB | 11d8215bce91a156e66c948134c725cc |
| ENCFF792WBY.tabix.bed.gz.dvc | 106.0 B | 5c13113fb8ec391d1bae1cac4d32d069 |
| ENCFF792WBY.tabix.bed.gz.tbi | 188.5 KB | 1e6c4fcf09c3edf39d6fcec02af8755f |
| ENCFF792WBY.tabix.bed.gz.tbi.dvc | 110.0 B | 04be7a746d60d08e2c182473f28123b3 |
| genomic_resource.yaml | 2.66 KB | 99d2f579d7da73bc3462e1489aa4f46d |
| genomic_resource_original.yaml | 2.4 KB | aa6c4d6fb9f10f5c3a19362901317759 |
| statistics/ |