DNase-seq_ENCSR550UWM
| Id: | DNase-seq/ENCSR550UWM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR550UWM [biosample_summary="Homo sapiens renal cortex interstitium tissue male embryo (97 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (97 days) output_type: peaks audit_warning: Alignment file {ENCFF190XEM|/files/ENCFF190XEM/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 21132751 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
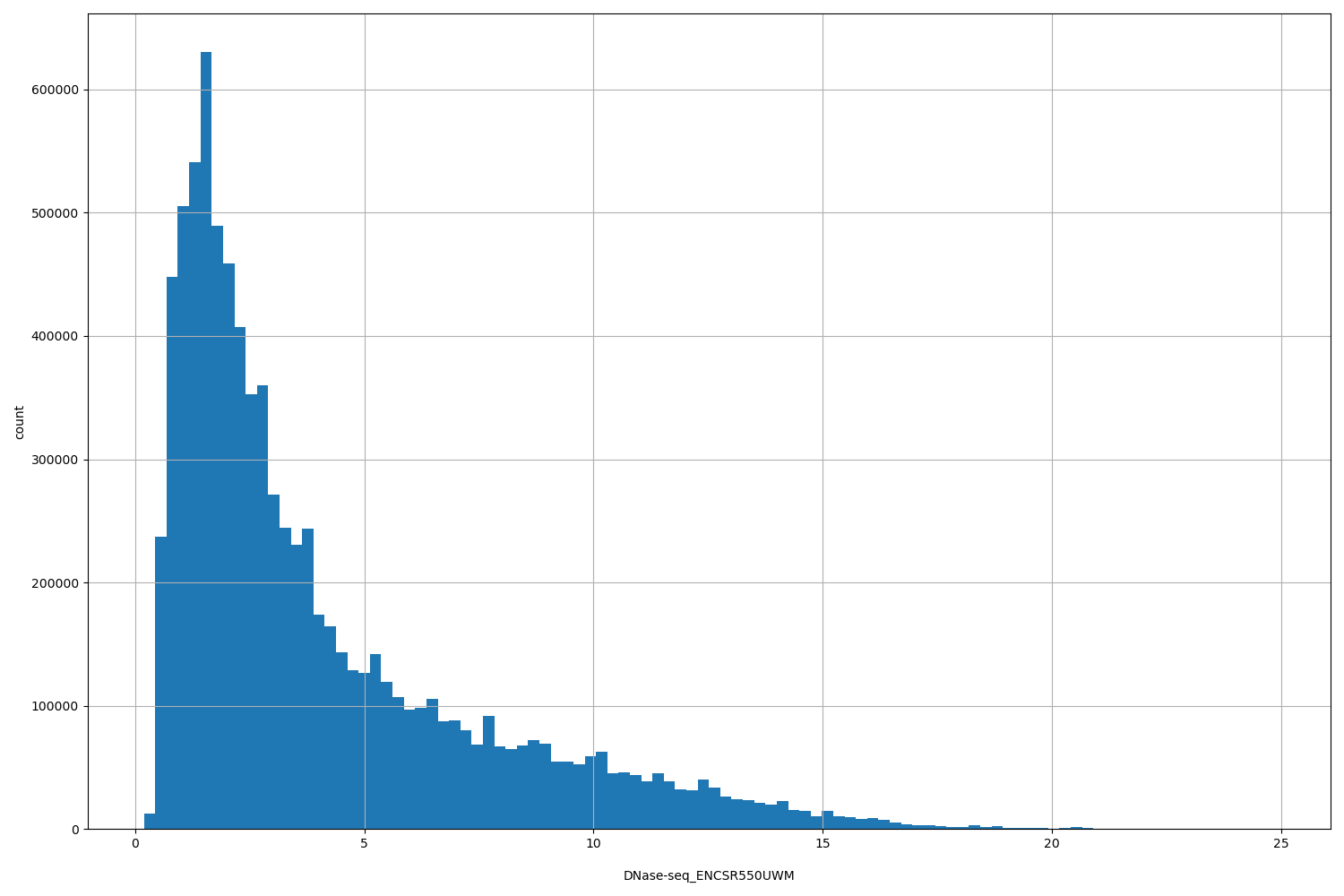
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR550UWM | float |
DNase-seq_ENCSR550UWM |
DNase-seq ENCSR550UWM [biosample_summary="Homo sapiens renal cortex interstitium tissue male embryo (97 days)"]
|
 |
[0.19, 24.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF912FDI.bed.gz | 487.87 KB | 32b220ad072d8df6c03823f937bdf1c2 |
| ENCFF912FDI.bed.gz.dvc | 100.0 B | d9eabf3e439eb6cabf9a0ac9091dd5a8 |
| ENCFF912FDI.tabix.bed.gz | 435.53 KB | dee6c32093a3d3954414ebeebf9598a6 |
| ENCFF912FDI.tabix.bed.gz.dvc | 106.0 B | c51934f19c49998e12dad9b5fe1cade6 |
| ENCFF912FDI.tabix.bed.gz.tbi | 259.45 KB | a7cbbda174062d5a575f6c9d01a83425 |
| ENCFF912FDI.tabix.bed.gz.tbi.dvc | 110.0 B | 03d20185577a1b3476c32718c9b7884d |
| genomic_resource.yaml | 1.87 KB | d8ff73c335fb9fffb322517911bc1a47 |
| genomic_resource_original.yaml | 1.76 KB | 77f8ea8a45f58e97bbb518e123abaada |
| statistics/ |