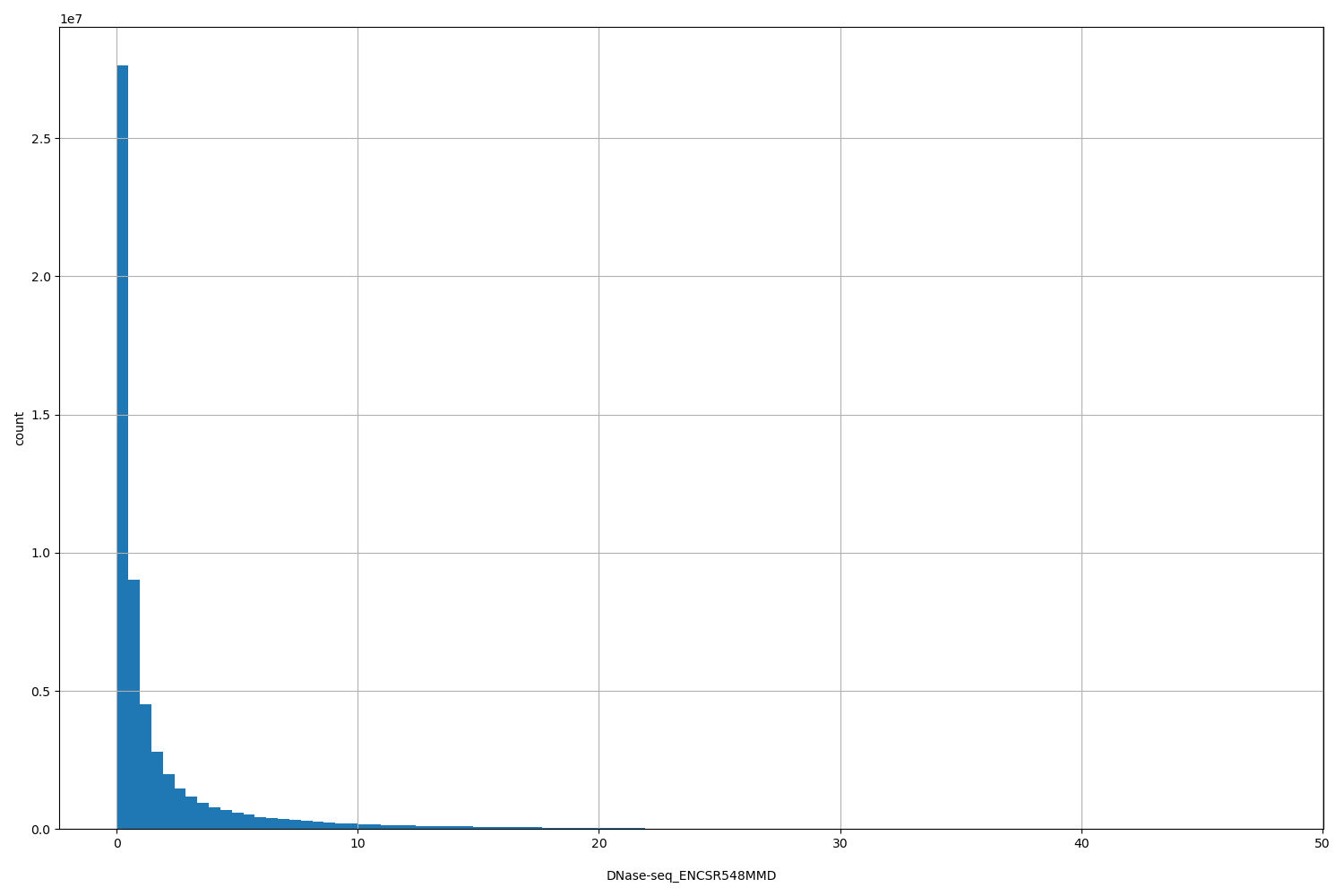
DNase-seq_ENCSR548MMD
| Id: | DNase-seq/ENCSR548MMD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR548MMD [biosample_summary="Homo sapiens EH"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Alignment file {ENCFF640FPX|/files/ENCFF640FPX/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 45505371 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR548MMD | float |
DNase-seq_ENCSR548MMD |
DNase-seq ENCSR548MMD [biosample_summary="Homo sapiens EH"]
|
 |
[0.0127, 47.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF628EXC.bed.gz | 3.22 MB | 8482014eec68574aa471373b062e857f |
| ENCFF628EXC.bed.gz.dvc | 101.0 B | 3b48f1ece2b3a3c69e1647b565f722a5 |
| ENCFF628EXC.tabix.bed.gz | 2.93 MB | 7d62ba94a9d864111aae2ed40872439d |
| ENCFF628EXC.tabix.bed.gz.dvc | 107.0 B | 71aa6179d55ac1f34840b28579f3d527 |
| ENCFF628EXC.tabix.bed.gz.tbi | 707.44 KB | 893e53a04bf98d2131699dc6624a6996 |
| ENCFF628EXC.tabix.bed.gz.tbi.dvc | 110.0 B | 17b1c162755b010c2d28e875efc5a96a |
| genomic_resource.yaml | 1.69 KB | 2acd07ee76c141ba0a7b0977fe977335 |
| genomic_resource_original.yaml | 1.62 KB | 139f44cbbe495972f817555a5c489430 |
| statistics/ |