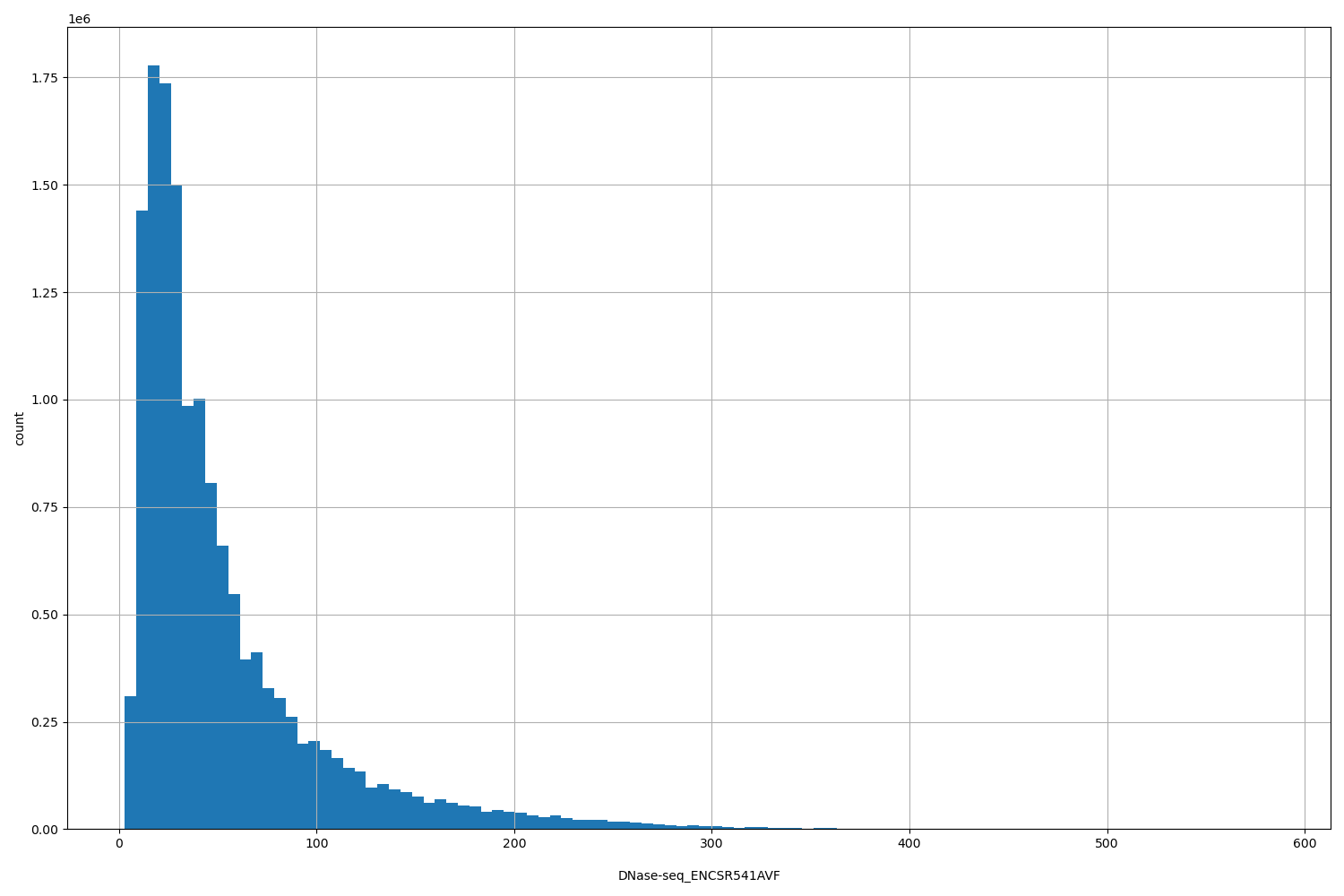
DNase-seq_ENCSR541AVF
| Id: | DNase-seq/ENCSR541AVF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR541AVF [biosample_summary="Homo sapiens left lung tissue female embryo (117 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (117 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN283BQK|/analyses/ENCAN283BQK/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF428XKG|/files/ENCFF428XKG/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 35679143 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF507MMD|/files/ENCFF507MMD/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 35443124 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR541AVF | float |
DNase-seq_ENCSR541AVF |
DNase-seq ENCSR541AVF [biosample_summary="Homo sapiens left lung tissue female embryo (117 days)"]
|
 |
[3, 584] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF690UKD.bed.gz | 952.4 KB | 6a03d7e55da2e497e10db0fc0b96f927 |
| ENCFF690UKD.bed.gz.dvc | 100.0 B | c5d48d3c1f53f45124fd72efd93da759 |
| ENCFF690UKD.tabix.bed.gz | 817.8 KB | 6b7a62047dd6d7f9a7365f61946f41f6 |
| ENCFF690UKD.tabix.bed.gz.dvc | 106.0 B | 82d28b97937908f2d3b7d3ec17e1370e |
| ENCFF690UKD.tabix.bed.gz.tbi | 473.29 KB | beed2d5abcd63beb06bf962807ebe515 |
| ENCFF690UKD.tabix.bed.gz.tbi.dvc | 110.0 B | ff12653b44d8c4e0e0afb0c26d9cd6f0 |
| genomic_resource.yaml | 2.59 KB | 3416dd7eef62dc1a7572db38130dd010 |
| genomic_resource_original.yaml | 2.5 KB | 7c077de2f54c4d7902c81c10a9cf91c6 |
| statistics/ |