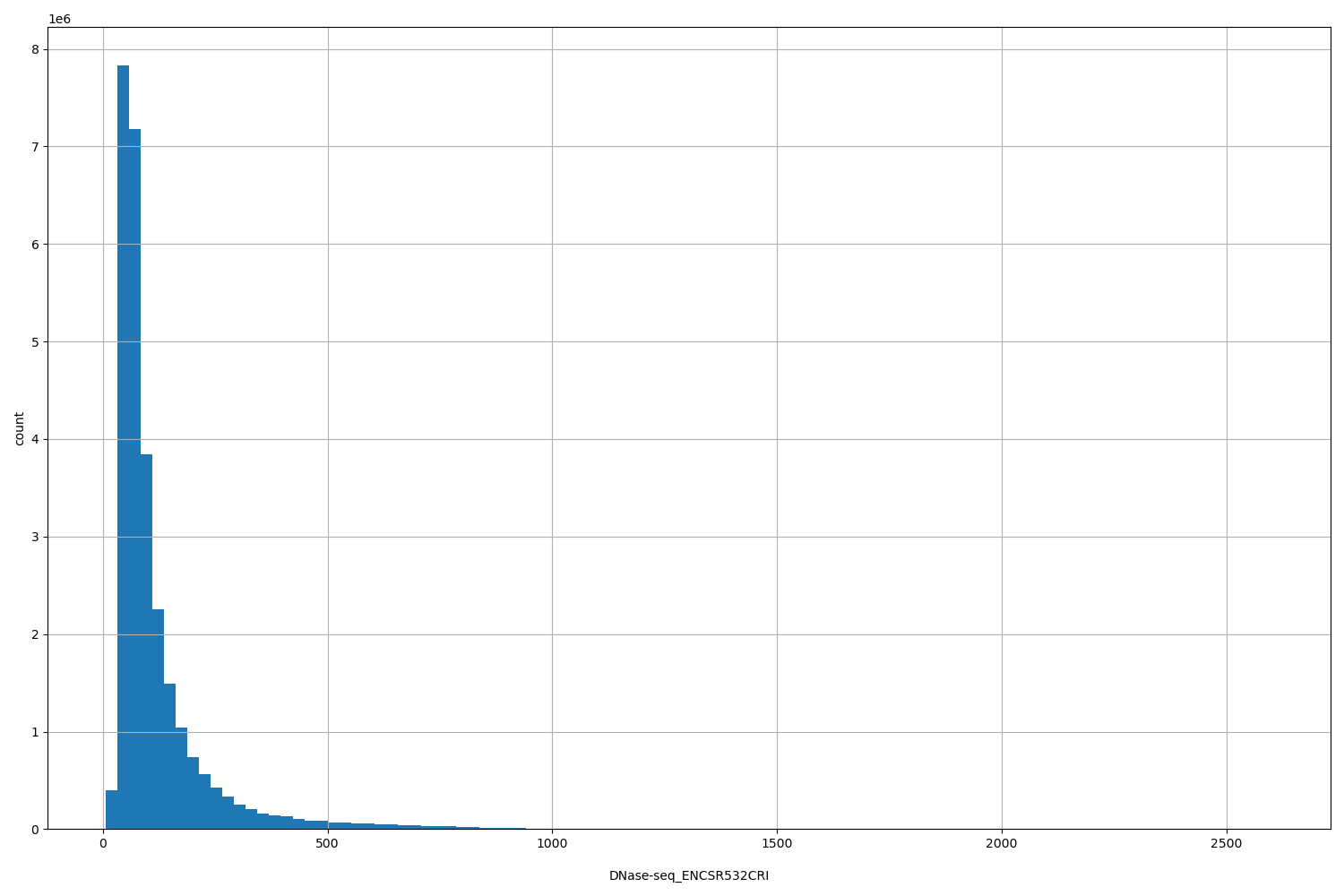
DNase-seq_ENCSR532CRI
| Id: | DNase-seq/ENCSR532CRI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR532CRI [biosample_summary="Homo sapiens limb tissue embryo (58 days) and embryo (59 days)"] |
| Description: |
status: released biological_replicates: Rep 2 summary: embryo (59 days) embryo (58 days) output_type: peaks audit_error: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF371ZLX|/files/ENCFF371ZLX/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL202DNS|/pipelines/ENCPL202DNS/}) for GRCh38 assembly have a SPOT1 score of 0.25. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_internal_action: Archived analysis {ENCAN929UJL|/analyses/ENCAN929UJL/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_not_compliant: Replicate concordance in DNase-seq experiments is measured by calculating the Pearson correlation between signal quantification of the replicates. ENCODE processed signal files {ENCFF850IPE|/files/ENCFF850IPE/}, {ENCFF135BMZ|/files/ENCFF135BMZ/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL202DNS|/pipelines/ENCPL202DNS/}) for GRCh38 assembly have a Pearson correlation of 0.83. According to ENCODE standards, in an anisogenic assay a Pearson correlation value > 0.85 is recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/} ) audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF757WSE|/files/ENCFF757WSE/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL202DNS|/pipelines/ENCPL202DNS/}) for GRCh38 assembly have a SPOT1 score of 0.25. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR532CRI | float |
DNase-seq_ENCSR532CRI |
DNase-seq ENCSR532CRI [biosample_summary="Homo sapiens limb tissue embryo (58 days) and embryo (59 days)"]
|
 |
[7, 2.6e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF551YAW.bed.gz | 1.78 MB | 88c48fefb709b5083fc4e9099f637348 |
| ENCFF551YAW.bed.gz.dvc | 101.0 B | 175ef7b39b722ca7ebbad91b83962aa7 |
| ENCFF551YAW.tabix.bed.gz | 1.52 MB | 241c4470e26788ab8ccf20913dbcd681 |
| ENCFF551YAW.tabix.bed.gz.dvc | 107.0 B | 68515965c9bf3770d2b94dd4f27fa722 |
| ENCFF551YAW.tabix.bed.gz.tbi | 661.29 KB | 63523992f93bc0699f85dfb929bab15e |
| ENCFF551YAW.tabix.bed.gz.tbi.dvc | 110.0 B | 375b448f8c8c9e8b08692667cf5f494b |
| genomic_resource.yaml | 3.64 KB | a78530093327cea837aa69182da17002 |
| genomic_resource_original.yaml | 3.54 KB | 4e920600aa5b0c56b41dd3753e43b16d |
| statistics/ |