DNase-seq_ENCSR531HPD
| Id: | DNase-seq/ENCSR531HPD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR531HPD [biosample_summary="Homo sapiens large intestine tissue male embryo (105 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (105 days) output_type: peaks audit_warning: Replicate 1_5 has no significant footprints detected. audit_warning: Alignment file {ENCFF504RZQ|/files/ENCFF504RZQ/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 26061617 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
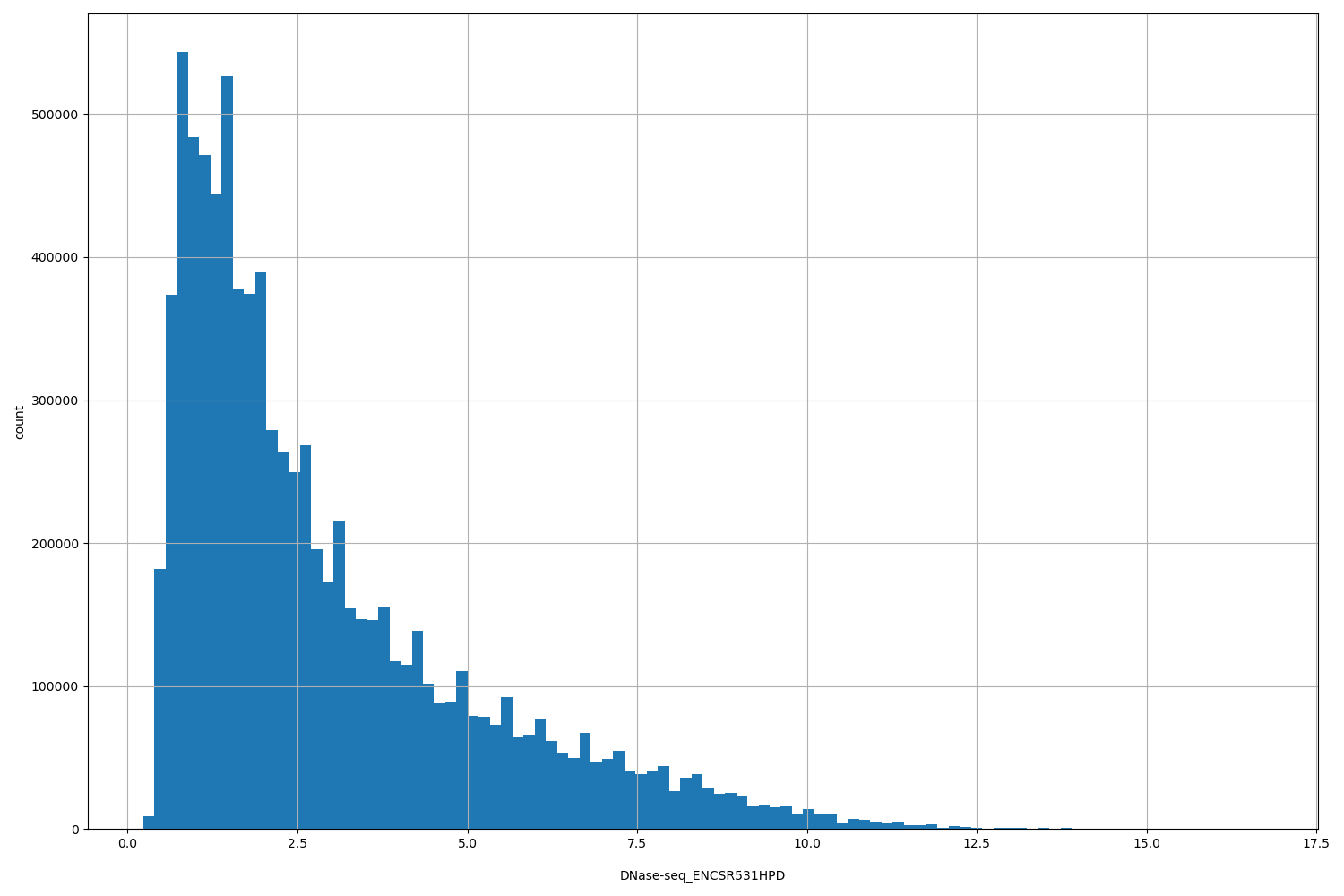
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR531HPD | float |
DNase-seq_ENCSR531HPD |
DNase-seq ENCSR531HPD [biosample_summary="Homo sapiens large intestine tissue male embryo (105 days)"]
|
 |
[0.231, 16.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF663QAD.bed.gz | 503.07 KB | e05ccebc6d1c806385bbfbef7d34e832 |
| ENCFF663QAD.bed.gz.dvc | 100.0 B | f39720fa6f4839eee9da839eb46239df |
| ENCFF663QAD.tabix.bed.gz | 448.23 KB | 8a51f1549fc38057154a768f4a594887 |
| ENCFF663QAD.tabix.bed.gz.dvc | 106.0 B | cbb46bf9fda3e88c72c52d5819fcf435 |
| ENCFF663QAD.tabix.bed.gz.tbi | 268.52 KB | d5c51320b5d51a5b20af7fffc6080737 |
| ENCFF663QAD.tabix.bed.gz.tbi.dvc | 110.0 B | b8389ab4d6fe222ab027d1c88cfbd0e3 |
| genomic_resource.yaml | 1.92 KB | 9e7e0d85e1c0b8096744cc8b4b05a168 |
| genomic_resource_original.yaml | 1.82 KB | be51631b423c3c67c26900585c2a3f81 |
| statistics/ |