DNase-seq_ENCSR488PHT
| Id: | DNase-seq/ENCSR488PHT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR488PHT [biosample_summary="Homo sapiens left lung tissue male embryo (113 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (113 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN758KMY|/analyses/ENCAN758KMY/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF062TCM|/files/ENCFF062TCM/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 38883695 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF737ILJ|/files/ENCFF737ILJ/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 38640691 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
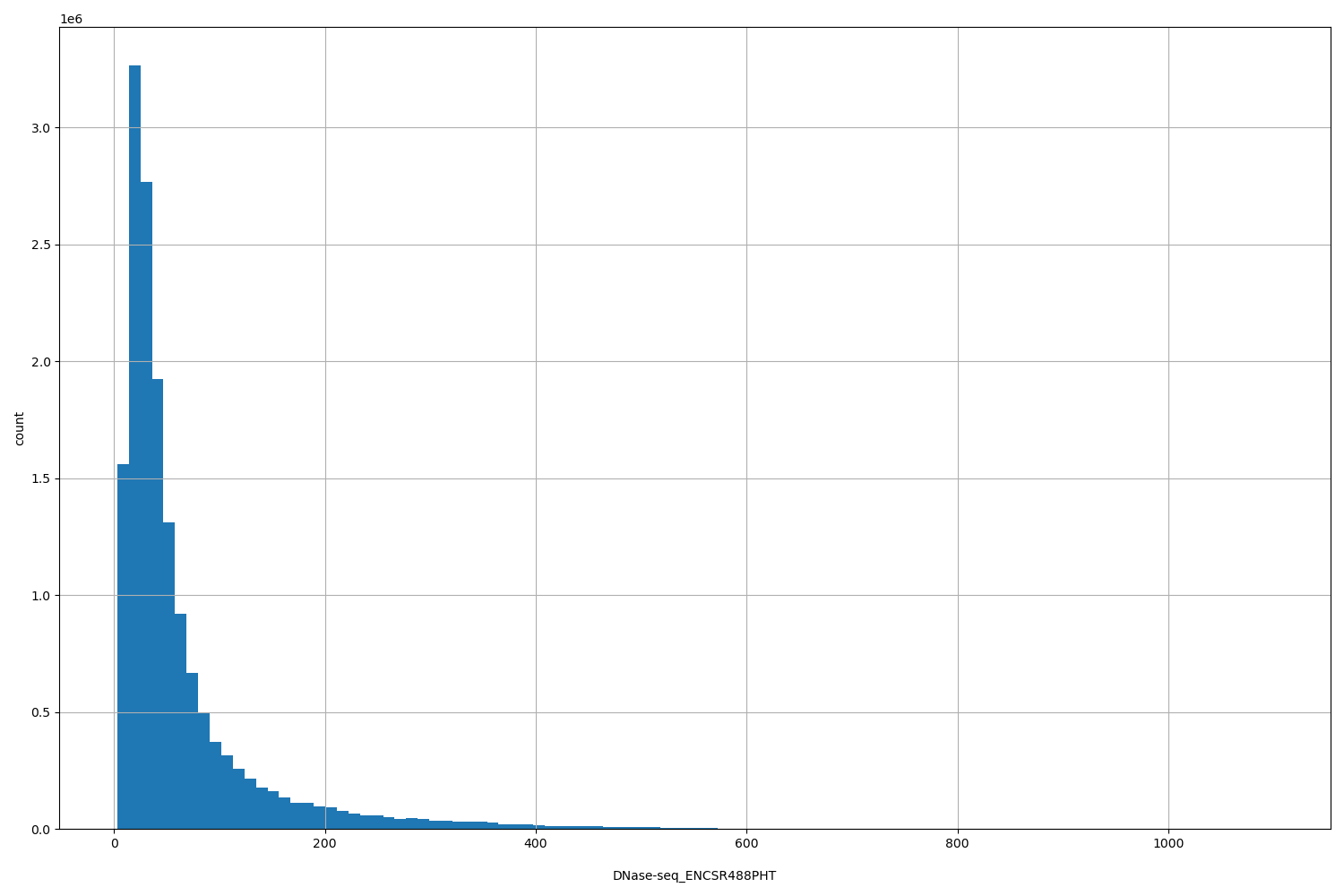
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR488PHT | float |
DNase-seq_ENCSR488PHT |
DNase-seq ENCSR488PHT [biosample_summary="Homo sapiens left lung tissue male embryo (113 days)"]
|
 |
[3, 1.1e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF676GRC.bed.gz | 1023.14 KB | 5154ab70fd7405835a6b0a822dffd6db |
| ENCFF676GRC.bed.gz.dvc | 101.0 B | 124a9245332fb6a10fa646c1436154ff |
| ENCFF676GRC.tabix.bed.gz | 879.37 KB | 0250c9c390ecead975c5ef548dabf880 |
| ENCFF676GRC.tabix.bed.gz.dvc | 106.0 B | 5be39fd0558a44c7a850efe4da0f7af7 |
| ENCFF676GRC.tabix.bed.gz.tbi | 484.08 KB | 97c7c58afcdcfd70b2e3a42c0801e612 |
| ENCFF676GRC.tabix.bed.gz.tbi.dvc | 110.0 B | 5f27fded09b8304e4721ae7432b3c0b9 |
| genomic_resource.yaml | 2.58 KB | 8ca36f25348dc18f6cea9ad6eb606fea |
| genomic_resource_original.yaml | 2.49 KB | 52142423ae877c96a41252f82ee6acc1 |
| statistics/ |