DNase-seq_ENCSR438TWI
| Id: | DNase-seq/ENCSR438TWI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR438TWI [biosample_summary="Homo sapiens muscle of back tissue male embryo (96 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (96 days) output_type: peaks audit_warning: Alignment file {ENCFF081IPQ|/files/ENCFF081IPQ/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 31153211 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
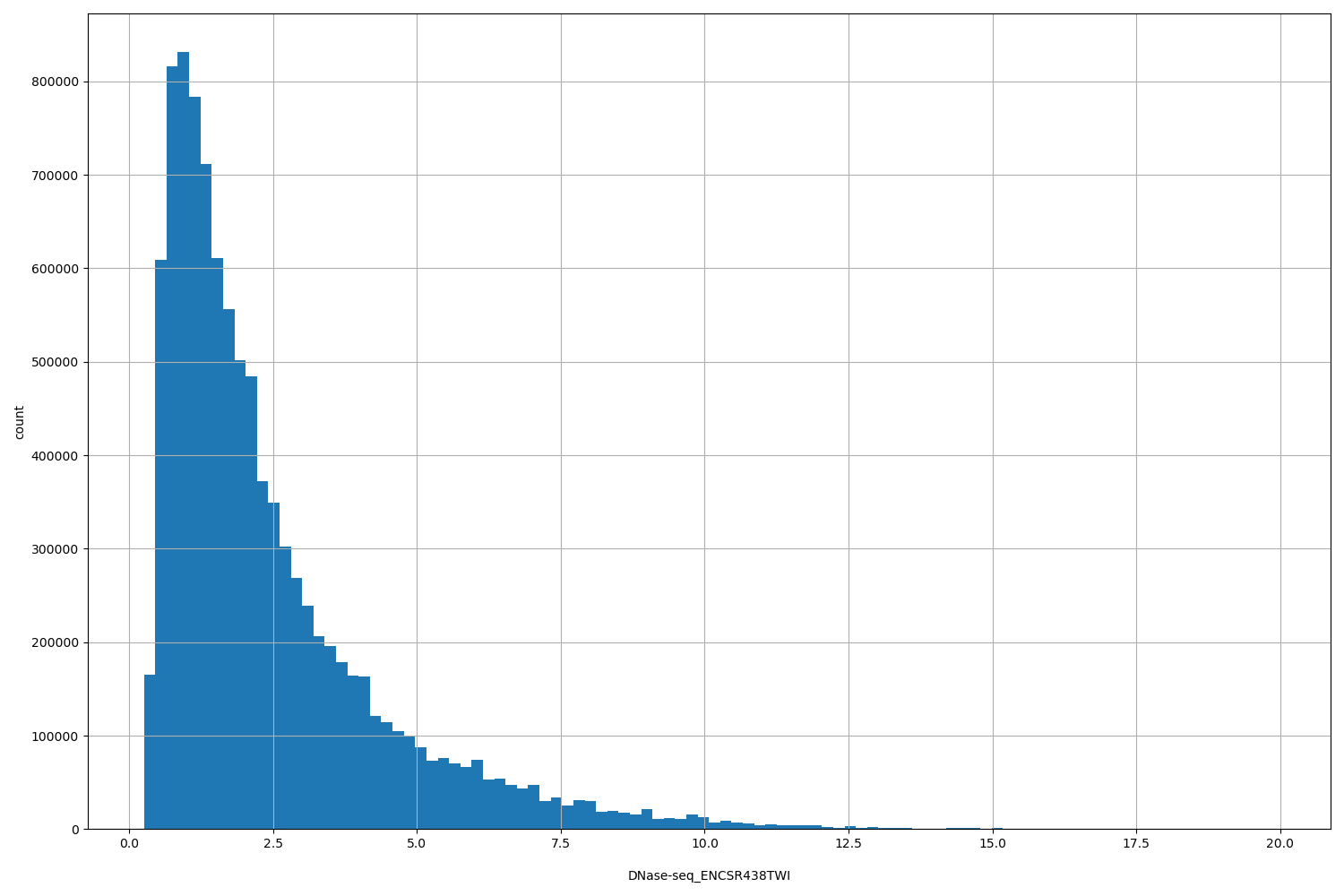
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR438TWI | float |
DNase-seq_ENCSR438TWI |
DNase-seq ENCSR438TWI [biosample_summary="Homo sapiens muscle of back tissue male embryo (96 days)"]
|
 |
[0.258, 19.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF378QIP.bed.gz | 542.18 KB | e17ccc1e0543d0788698e3b84089dd32 |
| ENCFF378QIP.bed.gz.dvc | 100.0 B | 3971d4ed4994ed50d4697bcf78129a5b |
| ENCFF378QIP.tabix.bed.gz | 486.79 KB | 9285928e2c4bb228c4fdc74df594bcf7 |
| ENCFF378QIP.tabix.bed.gz.dvc | 106.0 B | 783f818b84d066c170a26701a4d5b2f4 |
| ENCFF378QIP.tabix.bed.gz.tbi | 285.71 KB | 30af85aae1c80af195a2cedb6014b6bb |
| ENCFF378QIP.tabix.bed.gz.tbi.dvc | 110.0 B | 5fd733bd0f86a1c1963938ee3578bd44 |
| genomic_resource.yaml | 1.84 KB | ce31fcb57ede33df2943a165c06694f7 |
| genomic_resource_original.yaml | 1.73 KB | 48f148e84a01f96fb5232279b5a86d3c |
| statistics/ |