DNase-seq_ENCSR420NOA
| Id: | DNase-seq/ENCSR420NOA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR420NOA [biosample_summary="Homo sapiens hematopoietic multipotent progenitor cell treated with interleukin-3 for 11 days, kit ligand for 11 days, hydrocortisone succinate for 11 days, erythropoietin for 11 days"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (25 years) treated with interleukin-3 for 11 days, kit ligand for 11 days, hydrocortisone succinate for 11 days, erythropoietin for 11 days output_type: peaks audit_error: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF893TZO|/files/ENCFF893TZO/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.14. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF973FCP|/files/ENCFF973FCP/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.27. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF255ALH|/files/ENCFF255ALH/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.26. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
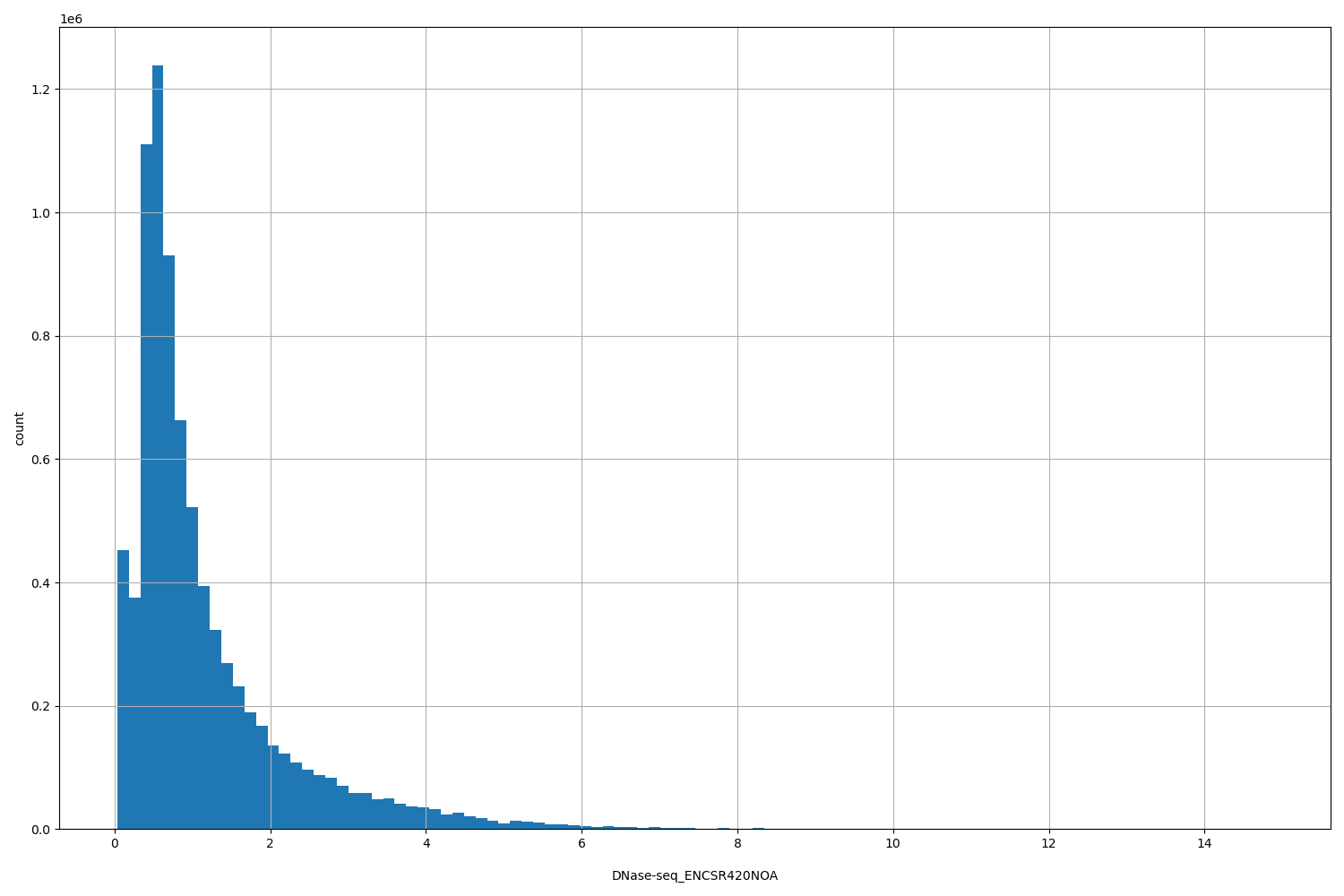
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR420NOA | float |
DNase-seq_ENCSR420NOA |
DNase-seq ENCSR420NOA [biosample_summary="Homo sapiens hematopoietic multipotent progenitor cell treated with interleukin-3 for 11 days, kit ligand for 11 days, hydrocortisone succinate for 11 days, erythropoietin for 11 days"]
|
 |
[0.0352, 14.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF377CYT.bed.gz | 452.91 KB | 66b14dc722ec42587d1408ebfb89c7b9 |
| ENCFF377CYT.bed.gz.dvc | 100.0 B | 67749a2602550ce2d9eb426ee443ae3d |
| ENCFF377CYT.tabix.bed.gz | 416.06 KB | dc6f05e5be55fb94fc742d6187d90b49 |
| ENCFF377CYT.tabix.bed.gz.dvc | 106.0 B | 7bff826e2b7174df7421c5fada6236fc |
| ENCFF377CYT.tabix.bed.gz.tbi | 213.88 KB | 643e71d63f7843232b54da83df1136e3 |
| ENCFF377CYT.tabix.bed.gz.tbi.dvc | 110.0 B | 6c95a2756ee961874c3a418295fa013b |
| genomic_resource.yaml | 4.36 KB | 6dc9a29ef2665c5b4739625083e8de9f |
| genomic_resource_original.yaml | 4.13 KB | d498dc85ccef24a51d4c2d0fa89f9df5 |
| statistics/ |