DNase-seq_ENCSR366EQH
| Id: | DNase-seq/ENCSR366EQH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR366EQH [biosample_summary="Homo sapiens large intestine tissue male embryo (115 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (115 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN262SVD|/analyses/ENCAN262SVD/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF163MHN|/files/ENCFF163MHN/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 22337770 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF258ARL|/files/ENCFF258ARL/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 22156276 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
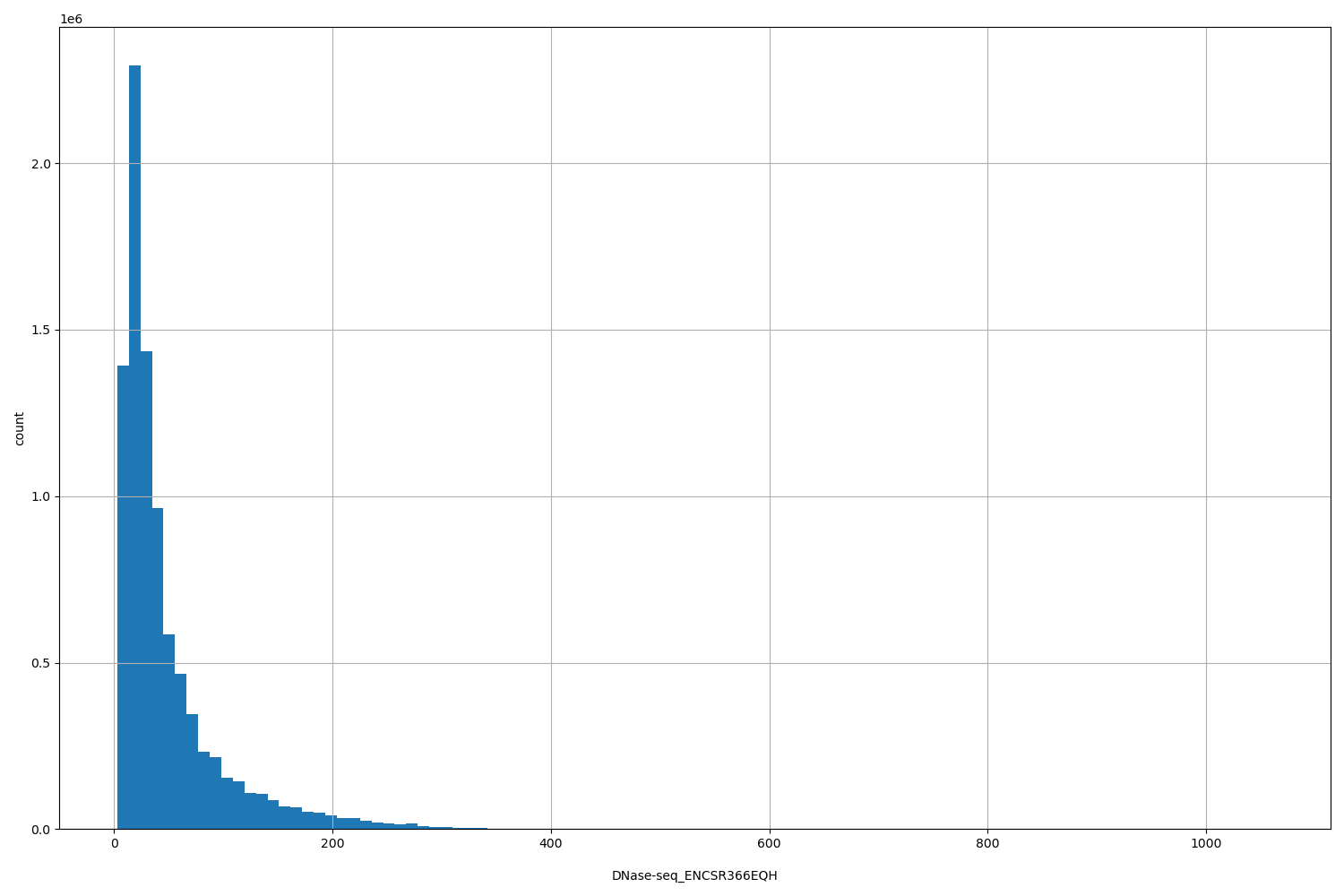
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR366EQH | float |
DNase-seq_ENCSR366EQH |
DNase-seq ENCSR366EQH [biosample_summary="Homo sapiens large intestine tissue male embryo (115 days)"]
|
 |
[3, 1.06e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF343JUT.bed.gz | 584.55 KB | b74551268c7ebfb23f194bdaafab0d49 |
| ENCFF343JUT.bed.gz.dvc | 100.0 B | b3b557fd93fb88c12ef33f6c073eb6bd |
| ENCFF343JUT.tabix.bed.gz | 500.5 KB | 569a978fd8d0151b1370650344354462 |
| ENCFF343JUT.tabix.bed.gz.dvc | 106.0 B | 14d9003f349682660ed872465b911634 |
| ENCFF343JUT.tabix.bed.gz.tbi | 322.38 KB | 82a710198a875013d85f53b5eb8857ed |
| ENCFF343JUT.tabix.bed.gz.tbi.dvc | 110.0 B | 2f6455d3de4bb8bed159085ae2778c57 |
| genomic_resource.yaml | 2.6 KB | a3996a6ccf5764023eb4f7b36eb56e2b |
| genomic_resource_original.yaml | 2.5 KB | 434bfcaa8c1404f1b45d304e6f4ce22b |
| statistics/ |