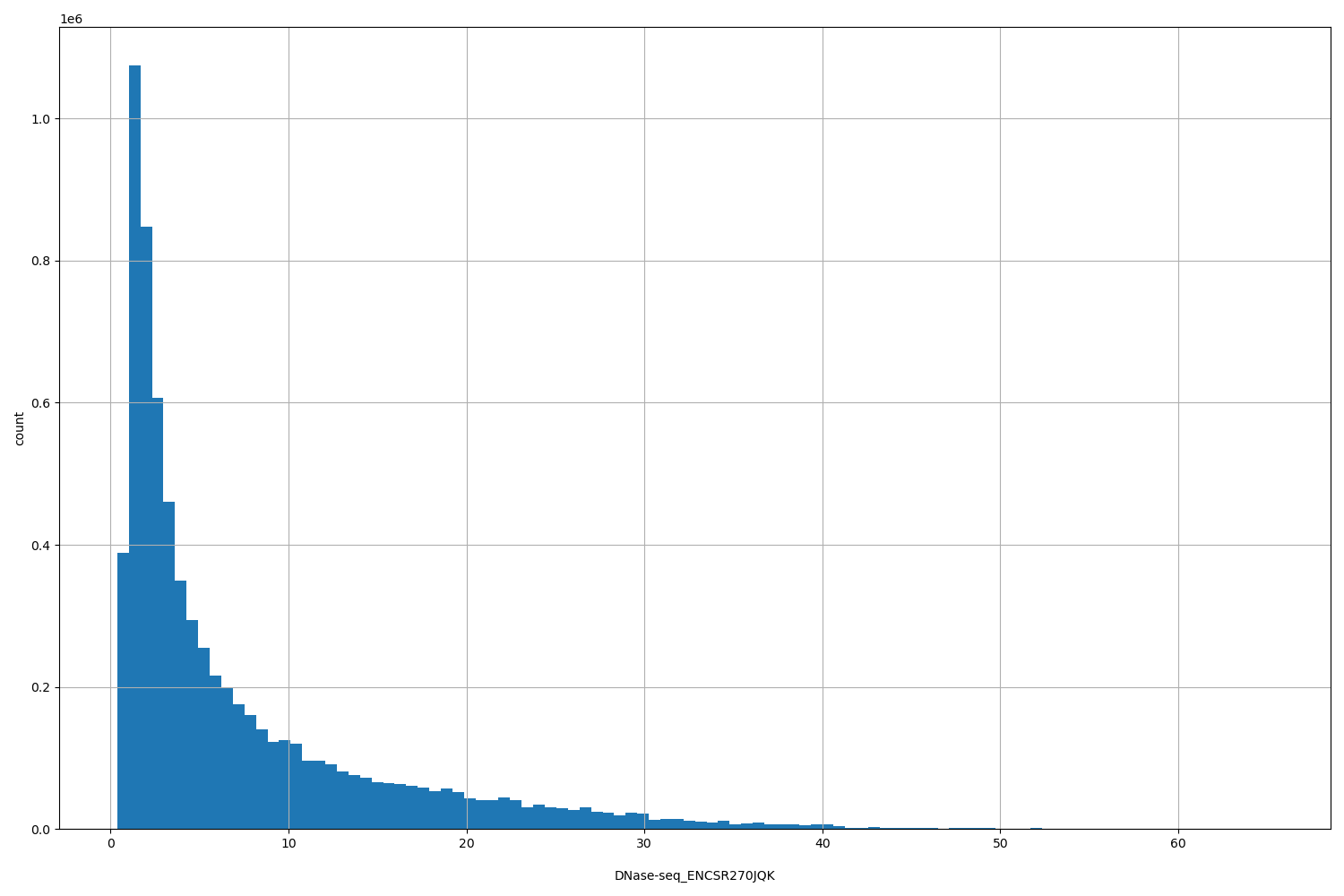
DNase-seq_ENCSR270JQK
| Id: | DNase-seq/ENCSR270JQK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR270JQK [biosample_summary="Homo sapiens T-cell female adult (40 years)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult (40 years) output_type: peaks audit_warning: Alignment file {ENCFF727ZRD|/files/ENCFF727ZRD/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 41836838 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR270JQK | float |
DNase-seq_ENCSR270JQK |
DNase-seq ENCSR270JQK [biosample_summary="Homo sapiens T-cell female adult (40 years)"]
|
 |
[0.383, 65.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF696CWC.bed.gz | 436.85 KB | ed239e505b11692b40af0b632738257a |
| ENCFF696CWC.bed.gz.dvc | 100.0 B | 9b194f62f0785116ecd3e2e955875df3 |
| ENCFF696CWC.tabix.bed.gz | 395.53 KB | 991de99e2e9e3e043ee391c538107633 |
| ENCFF696CWC.tabix.bed.gz.dvc | 106.0 B | 97b32bf4972df8a085f4f387f243bb09 |
| ENCFF696CWC.tabix.bed.gz.tbi | 201.84 KB | 7414fbbd813e52255dcbc45a1da62f71 |
| ENCFF696CWC.tabix.bed.gz.tbi.dvc | 110.0 B | ae91324693bf2c5ec707c895f6b0c05c |
| genomic_resource.yaml | 1.8 KB | 14b534b216bbc89408797f65272716c4 |
| genomic_resource_original.yaml | 1.71 KB | 4f2d5318d94a4bf3acfb0fee916fbc58 |
| statistics/ |