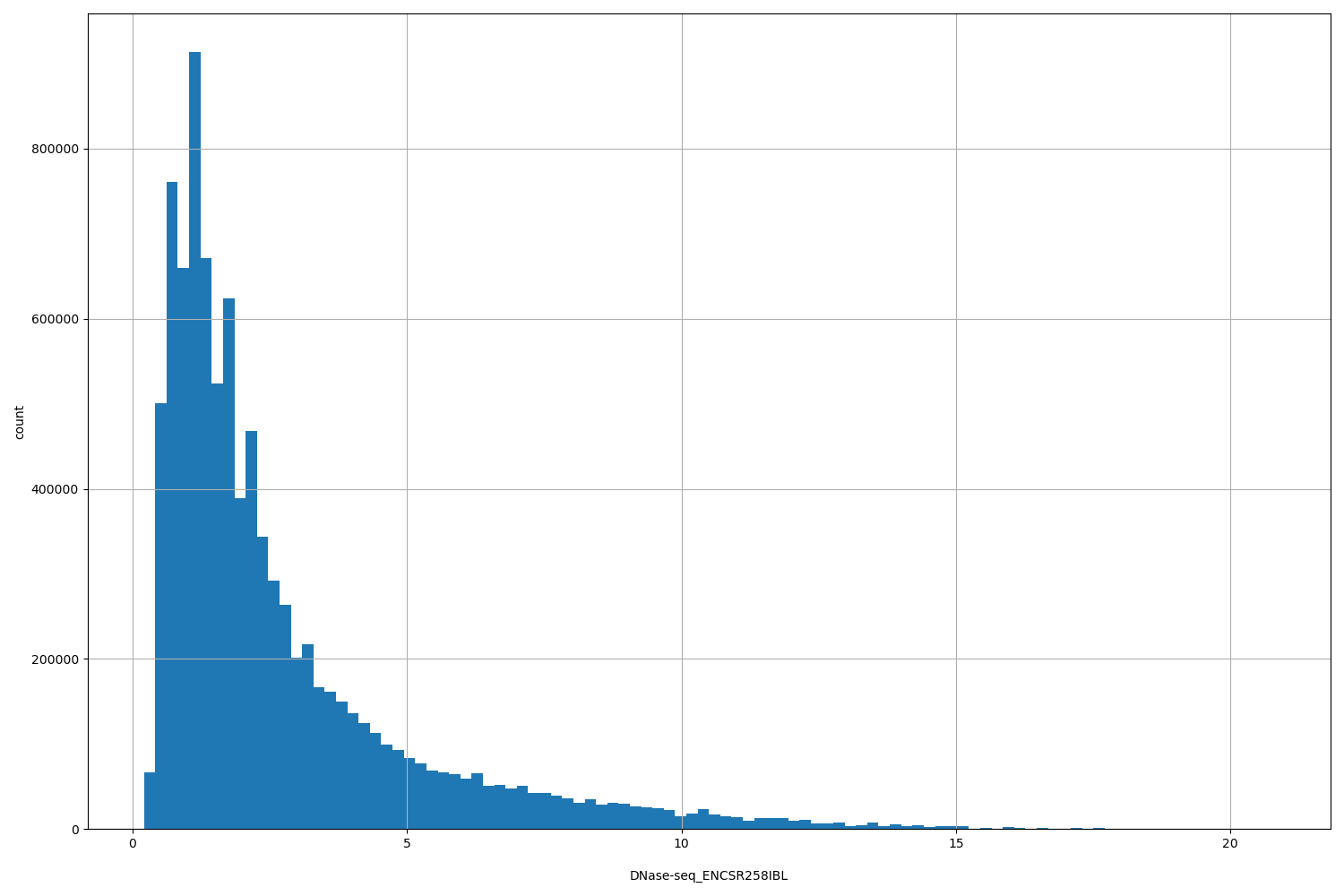
DNase-seq_ENCSR258IBL
| Id: | DNase-seq/ENCSR258IBL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR258IBL [biosample_summary="Homo sapiens left renal pelvis tissue male embryo (105 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (105 days) output_type: peaks audit_warning: Alignment file {ENCFF268QVD|/files/ENCFF268QVD/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 29225950 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR258IBL | float |
DNase-seq_ENCSR258IBL |
DNase-seq ENCSR258IBL [biosample_summary="Homo sapiens left renal pelvis tissue male embryo (105 days)"]
|
 |
[0.206, 20.8] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF322IFH.bed.gz | 551.06 KB | 04f52fe402a5294f8b055740c21e93c4 |
| ENCFF322IFH.bed.gz.dvc | 100.0 B | 1bfc0a993f84e55e719a1b8872ebf642 |
| ENCFF322IFH.tabix.bed.gz | 489.16 KB | d2183c01bb64e9b2eda97cb1c4eaadf9 |
| ENCFF322IFH.tabix.bed.gz.dvc | 106.0 B | 38d643e5775d4bbba70c2c5f536bfd7b |
| ENCFF322IFH.tabix.bed.gz.tbi | 296.7 KB | cedcca37818da3809c91b31fa447fcbf |
| ENCFF322IFH.tabix.bed.gz.tbi.dvc | 110.0 B | 09948bacbd7ef7c672e34c4060806765 |
| genomic_resource.yaml | 1.85 KB | 897e5f8171bdaf4a039a8c7d1387144d |
| genomic_resource_original.yaml | 1.74 KB | 2673af2c36ea24be1cb7cc0087d87e1a |
| statistics/ |