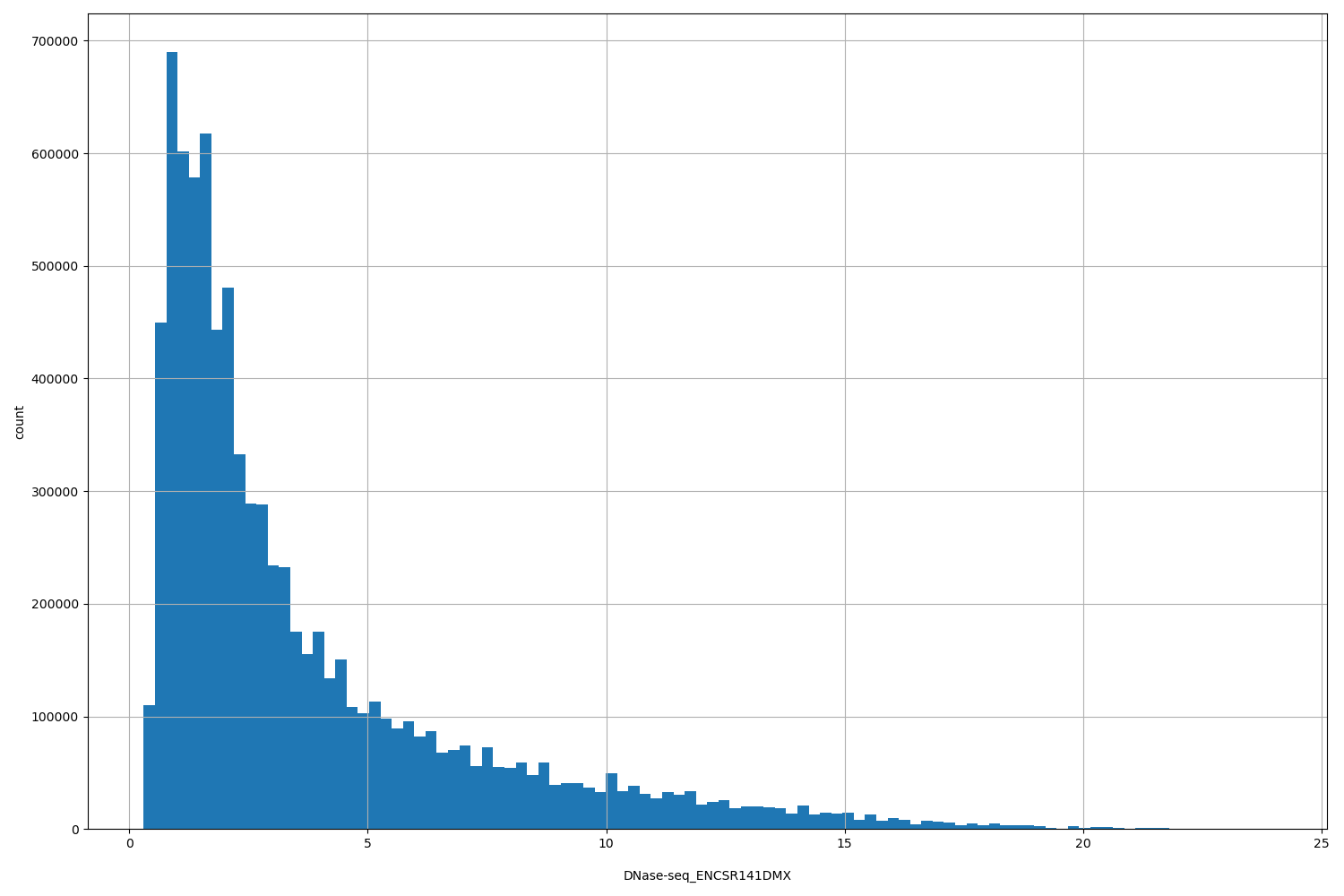
DNase-seq_ENCSR141DMX
| Id: | DNase-seq/ENCSR141DMX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR141DMX [biosample_summary="Homo sapiens renal pelvis tissue female embryo (96 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (96 days) output_type: peaks audit_warning: Alignment file {ENCFF777NPC|/files/ENCFF777NPC/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 22872720 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR141DMX | float |
DNase-seq_ENCSR141DMX |
DNase-seq ENCSR141DMX [biosample_summary="Homo sapiens renal pelvis tissue female embryo (96 days)"]
|
 |
[0.307, 23.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF166OIS.bed.gz | 489.96 KB | a743cbba77dbb124726df73cf857797d |
| ENCFF166OIS.bed.gz.dvc | 100.0 B | 324e6be5c7592516a9ea4b7989efb2ae |
| ENCFF166OIS.tabix.bed.gz | 435.61 KB | 45207f75c2238721aff67766b948bebb |
| ENCFF166OIS.tabix.bed.gz.dvc | 106.0 B | 6cb28a5150a7c68cd4121902b7d28008 |
| ENCFF166OIS.tabix.bed.gz.tbi | 265.25 KB | 18d67934c43dcaf146a01d9654a42ba7 |
| ENCFF166OIS.tabix.bed.gz.tbi.dvc | 110.0 B | b5466e0add07b6be2d64cde568e835e1 |
| genomic_resource.yaml | 1.84 KB | 295fbb410ca1e9f51b762a4f30241621 |
| genomic_resource_original.yaml | 1.74 KB | ed525a61a6720ebd721060c6ffac3805 |
| statistics/ |