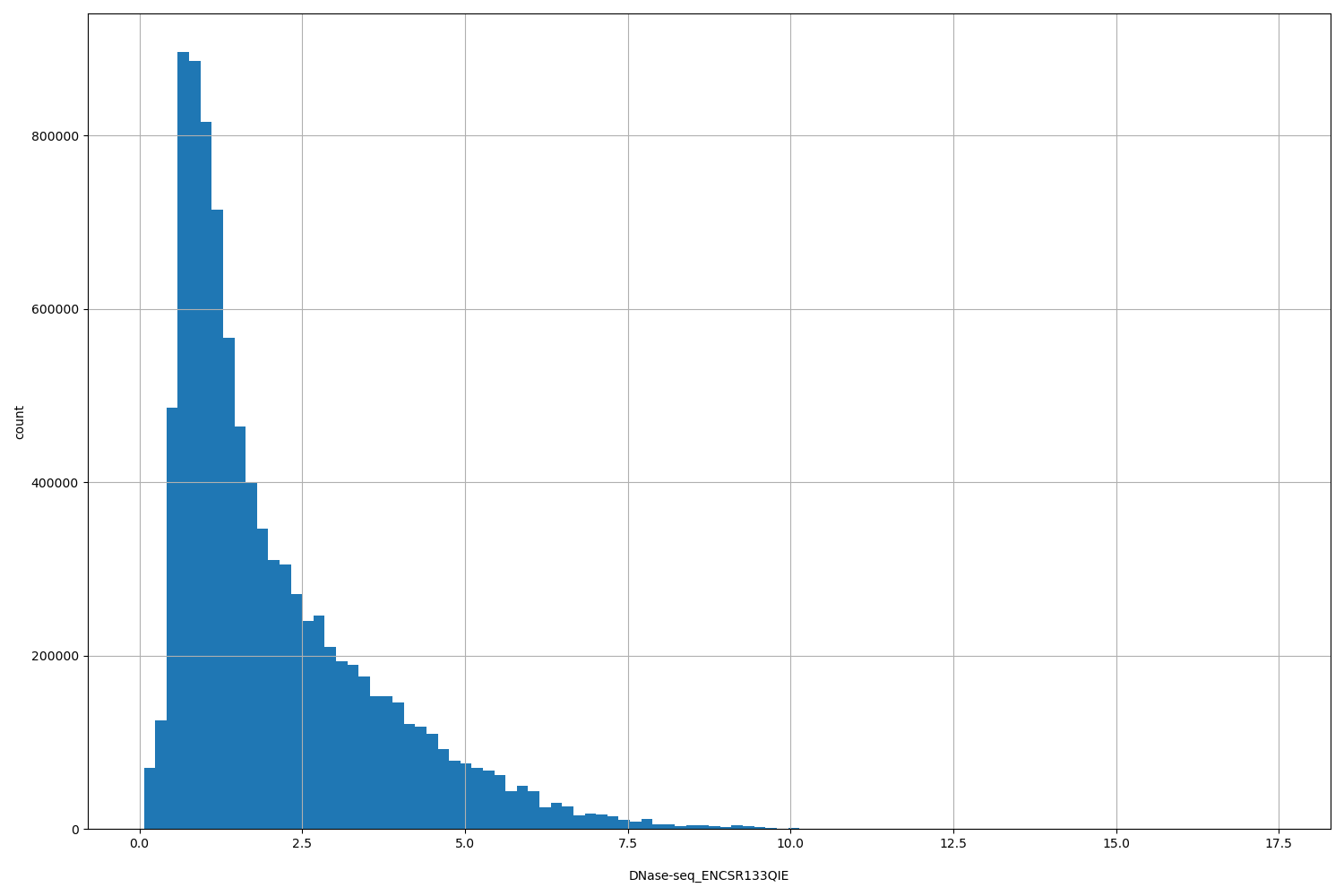
DNase-seq_ENCSR133QIE
| Id: | DNase-seq/ENCSR133QIE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR133QIE [biosample_summary="Homo sapiens stimulated activated effector memory CD8-positive, alpha-beta T cell male adult (33 years) treated with 100 ng/mL Interleukin-12 subunit alpha for 72 hours, anti-CD3 and anti-CD28 coated beads for 72 hours, 100 ng/mL Interleukin-12 subunit beta for 72 hours"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (33 years) treated with 100 ng/mL Interleukin-12 subunit alpha for 72 hours, anti-CD3 and anti-CD28 coated beads for 72 hours, 100 ng/mL Interleukin-12 subunit beta for 72 hours output_type: peaks audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF192RAV|/files/ENCFF192RAV/} processed by DNase-seq ENCODE4 v3.0.0 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.29. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR133QIE | float |
DNase-seq_ENCSR133QIE |
DNase-seq ENCSR133QIE [biosample_summary="Homo sapiens stimulated activated effector memory CD8-positive, alpha-beta T cell male adult (33 years) treated with 100 ng/mL Interleukin-12 subunit alpha for 72 hours, anti-CD3 and anti-CD28 coated beads for 72 hours, 100 ng/mL Interleukin-12 subunit beta for 72 hours"]
|
 |
[0.0683, 17.4] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF257BSS.bed.gz | 463.25 KB | 26dfb64af72555225defe5373ed9dba1 |
| ENCFF257BSS.bed.gz.dvc | 100.0 B | 19c9e8609f84da5fe87ec2f59503d78e |
| ENCFF257BSS.tabix.bed.gz | 429.41 KB | 4a2553418bd6ca196a0ed6d737e149e8 |
| ENCFF257BSS.tabix.bed.gz.dvc | 106.0 B | 7dc7a8794d32ad32dfa2ea46c44e1ee1 |
| ENCFF257BSS.tabix.bed.gz.tbi | 213.25 KB | 4ed5c9079ed7bc5b5cd71a012a078f6f |
| ENCFF257BSS.tabix.bed.gz.tbi.dvc | 110.0 B | cbff78f939643da7893a1f808b44ce5f |
| genomic_resource.yaml | 3.02 KB | 4c44cb834164f06ba546cff3b528c44d |
| genomic_resource_original.yaml | 2.72 KB | 5afa8d328d11a693ef8e071df68e13df |
| statistics/ |