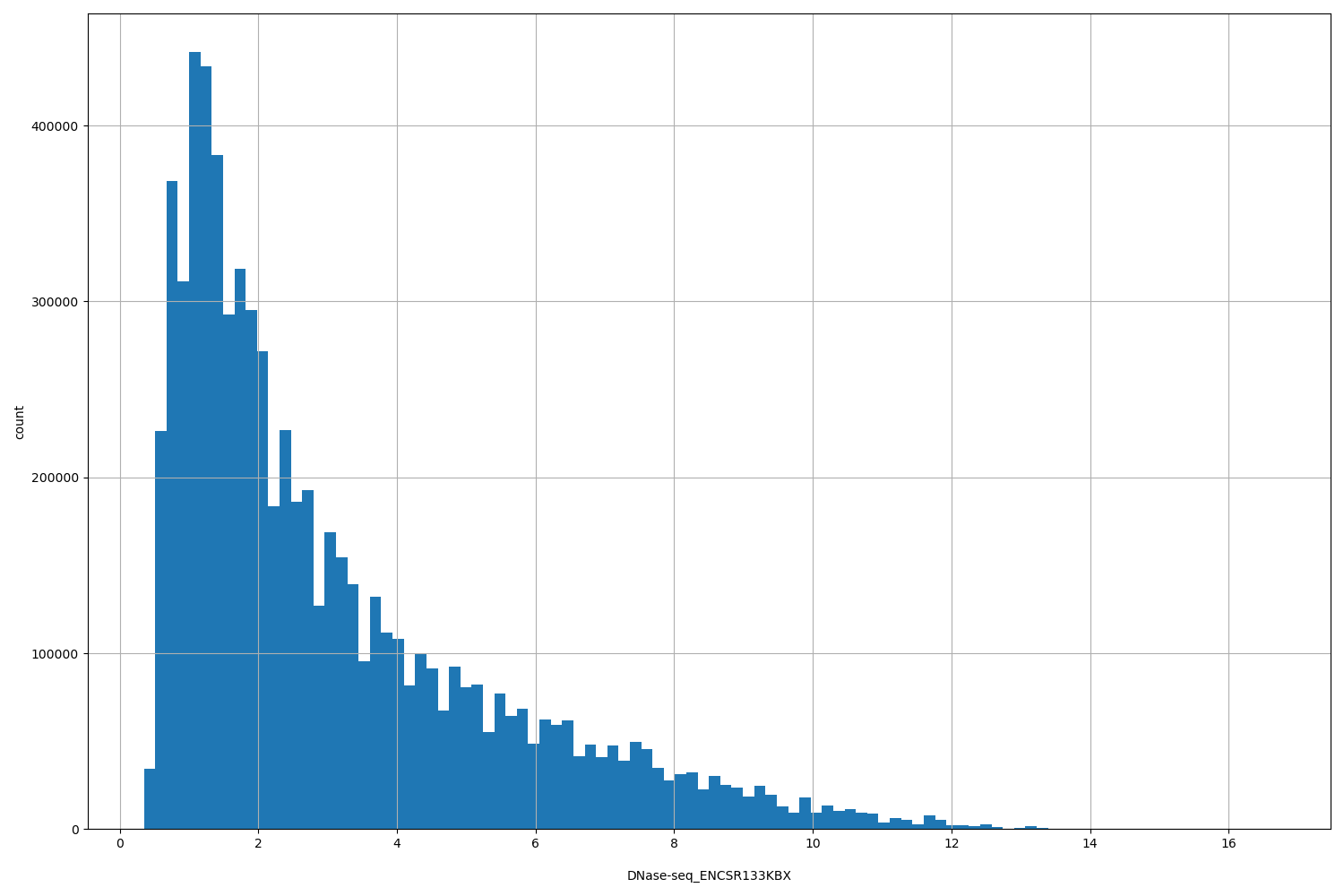
DNase-seq_ENCSR133KBX
| Id: | DNase-seq/ENCSR133KBX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR133KBX [biosample_summary="Homo sapiens small intestine tissue female embryo (107 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (107 days) output_type: peaks audit_warning: Alignment file {ENCFF753AOO|/files/ENCFF753AOO/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 23001916 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR133KBX | float |
DNase-seq_ENCSR133KBX |
DNase-seq ENCSR133KBX [biosample_summary="Homo sapiens small intestine tissue female embryo (107 days)"]
|
 |
[0.349, 16.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF540NZQ.bed.gz | 408.15 KB | 171cb3cef36413b722a2e953facc030f |
| ENCFF540NZQ.bed.gz.dvc | 100.0 B | 01d7875443954cc308f2ac7a4e7cb30d |
| ENCFF540NZQ.tabix.bed.gz | 361.7 KB | 9bb47f6317a626a82fa44148506d87c8 |
| ENCFF540NZQ.tabix.bed.gz.dvc | 106.0 B | 7d9a10becd90d1dc2ec420b116bf3991 |
| ENCFF540NZQ.tabix.bed.gz.tbi | 230.51 KB | 582fd72985b9e4a431a0d55019f235fc |
| ENCFF540NZQ.tabix.bed.gz.tbi.dvc | 110.0 B | 311c8770c13d7e9b17cadb68ba5169ef |
| genomic_resource.yaml | 1.85 KB | aca1bac3a7551d4b7e9836d3dab312f2 |
| genomic_resource_original.yaml | 1.75 KB | 2ebcf7727d6e959d6b6c4f9bf93de8db |
| statistics/ |