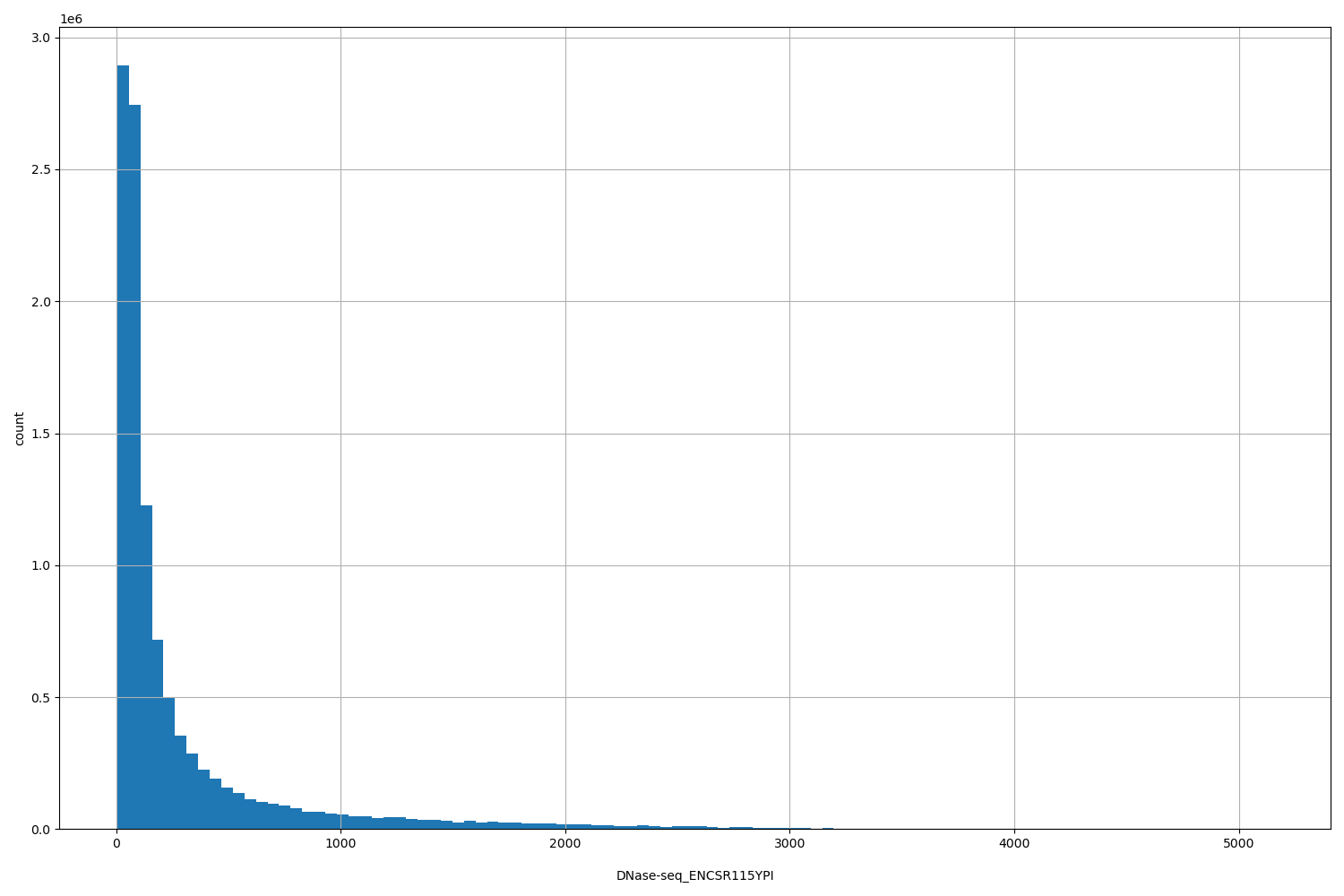
DNase-seq_ENCSR115YPI
| Id: | DNase-seq/ENCSR115YPI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR115YPI [biosample_summary="Homo sapiens hematopoietic multipotent progenitor cell treated with erythropoietin for 20 days, hydrocortisone succinate for 20 days, kit ligand for 20 days, interleukin-3 for 20 days"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male adult (25 years) treated with erythropoietin for 20 days, hydrocortisone succinate for 20 days, kit ligand for 20 days, interleukin-3 for 20 days output_type: peaks audit_error: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF986NWU|/files/ENCFF986NWU/} processed by DNase-seq ENCODE3 GRCh38 pipeline ({ENCPL202DNS|/pipelines/ENCPL202DNS/}) for GRCh38 assembly have a SPOT1 score of 0.22. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_internal_action: Archived analysis {ENCAN637UOT|/analyses/ENCAN637UOT/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR115YPI | float |
DNase-seq_ENCSR115YPI |
DNase-seq ENCSR115YPI [biosample_summary="Homo sapiens hematopoietic multipotent progenitor cell treated with erythropoietin for 20 days, hydrocortisone succinate for 20 days, kit ligand for 20 days, interleukin-3 for 20 days"]
|
 |
[5, 5.15e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF515CNP.bed.gz | 729.48 KB | e666868df6d1a64ab878d0e746cdff4a |
| ENCFF515CNP.bed.gz.dvc | 100.0 B | d697b11c0d49cac1cc84dbf73e0dc50e |
| ENCFF515CNP.tabix.bed.gz | 629.67 KB | 2f4dd4054a8b47c980b1c8e9d2445e6f |
| ENCFF515CNP.tabix.bed.gz.dvc | 106.0 B | b1f95e9d66203effd8a50bb51a3fe887 |
| ENCFF515CNP.tabix.bed.gz.tbi | 281.34 KB | a80295cf5ac480d06f8a2c803ad01ae9 |
| ENCFF515CNP.tabix.bed.gz.tbi.dvc | 110.0 B | 7d95783ba1a2eb8aec674f15f011ab79 |
| genomic_resource.yaml | 2.87 KB | a36cc7b54832ec213ef712ef04ef1795 |
| genomic_resource_original.yaml | 2.65 KB | a15f68c81e99a1a36d2d01786f4f6f14 |
| statistics/ |