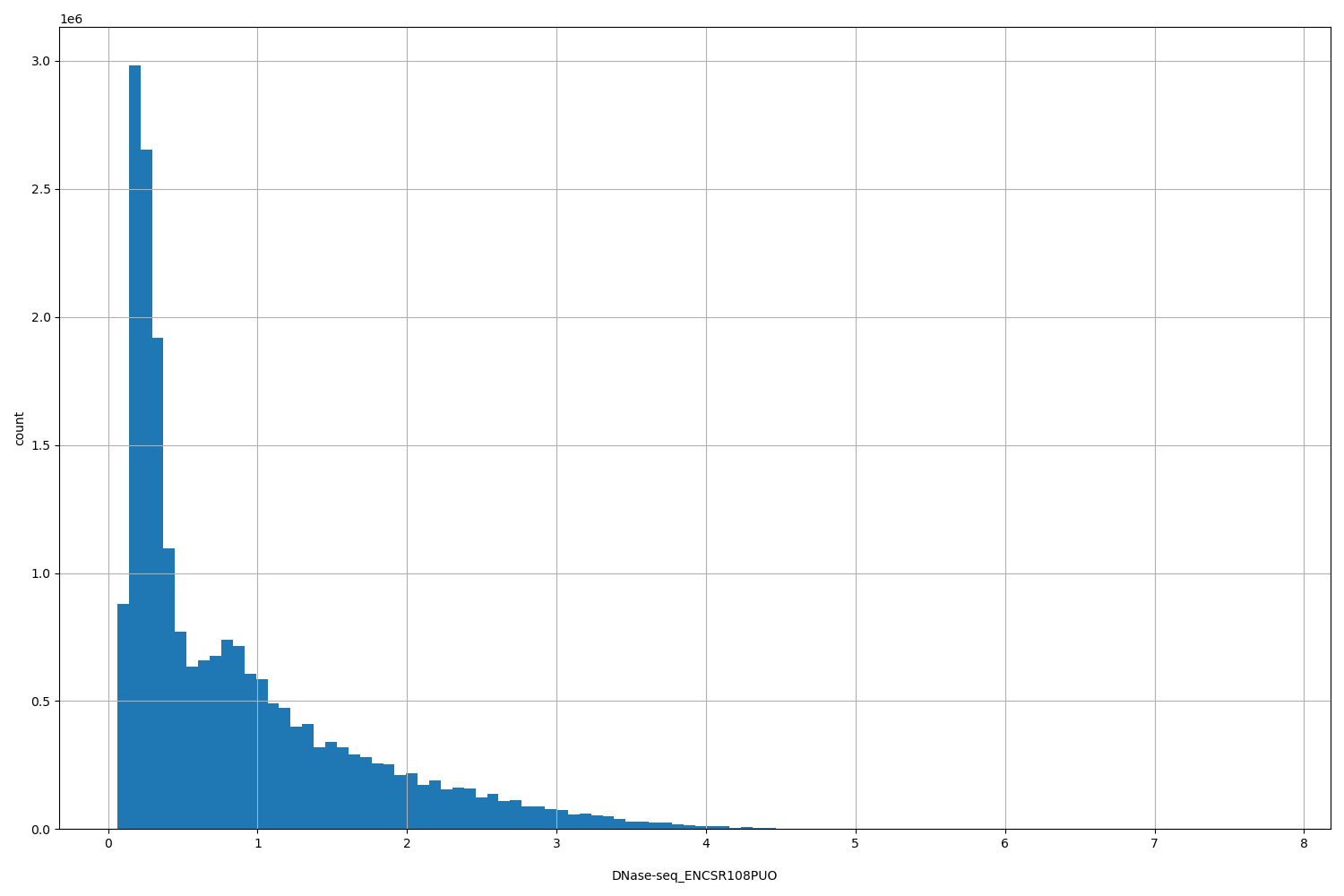
DNase-seq_ENCSR108PUO
| Id: | DNase-seq/ENCSR108PUO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR108PUO [biosample_summary="Homo sapiens GM23338"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF376XJI|/files/ENCFF376XJI/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.35. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR108PUO | float |
DNase-seq_ENCSR108PUO |
DNase-seq ENCSR108PUO [biosample_summary="Homo sapiens GM23338"]
|
 |
[0.0616, 7.79] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF580UGT.bed.gz | 935.79 KB | 2f5c0a3d032cb9ff70c9ca40046b551f |
| ENCFF580UGT.bed.gz.dvc | 100.0 B | 86f47560ea8574e0d83cb8b0d2c91a0d |
| ENCFF580UGT.tabix.bed.gz | 891.3 KB | bdb95fe3b36459cd1a97bc0cd546d44e |
| ENCFF580UGT.tabix.bed.gz.dvc | 106.0 B | 76178c2b217765d15e3b1fe8d4177d53 |
| ENCFF580UGT.tabix.bed.gz.tbi | 247.58 KB | a3ee757b86f18c8a3a695de66341b7fc |
| ENCFF580UGT.tabix.bed.gz.tbi.dvc | 110.0 B | 632a3a34900fafc0fc966a8340960db8 |
| genomic_resource.yaml | 1.92 KB | 65aa13d9af3a66e4aafdbc8263789809 |
| genomic_resource_original.yaml | 1.85 KB | 35eca779cf9c197f5fbc388a5b9514e5 |
| statistics/ |