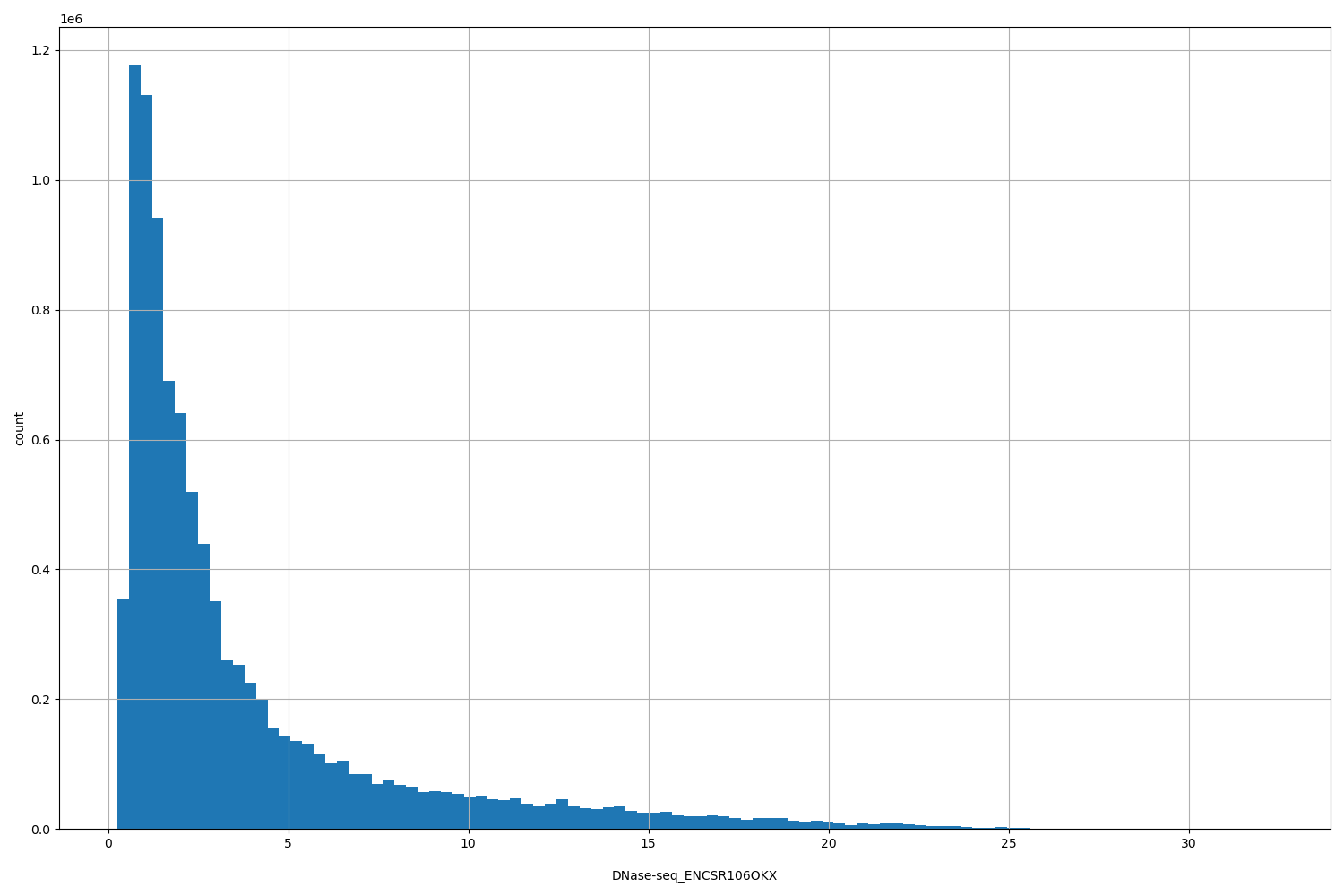
DNase-seq_ENCSR106OKX
| Id: | DNase-seq/ENCSR106OKX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR106OKX [biosample_summary="Homo sapiens kidney tissue female embryo (120 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (120 days) output_type: peaks audit_warning: Alignment file {ENCFF962IWA|/files/ENCFF962IWA/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 27450670 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR106OKX | float |
DNase-seq_ENCSR106OKX |
DNase-seq ENCSR106OKX [biosample_summary="Homo sapiens kidney tissue female embryo (120 days)"]
|
 |
[0.256, 32.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF388NNQ.bed.gz | 601.33 KB | 53d093fdc8a9603386b8e69f4a7aa71f |
| ENCFF388NNQ.bed.gz.dvc | 100.0 B | 63d459d0c5d853c6e86247bd40f91b0b |
| ENCFF388NNQ.tabix.bed.gz | 536.87 KB | dd6de313cb76beaef477405a5f2bd867 |
| ENCFF388NNQ.tabix.bed.gz.dvc | 106.0 B | 63cfd11ef60d854c65a9edc353b65b0e |
| ENCFF388NNQ.tabix.bed.gz.tbi | 313.95 KB | 1f561c10e39c1acaa3e663eef6bf1e81 |
| ENCFF388NNQ.tabix.bed.gz.tbi.dvc | 110.0 B | c5bd2d591676f970694ae33898dac2cd |
| genomic_resource.yaml | 1.83 KB | d112b0dd0d8e071eba19c9d9a5b54892 |
| genomic_resource_original.yaml | 1.73 KB | 561c5f0a40bf613b92d74303613deb3c |
| statistics/ |