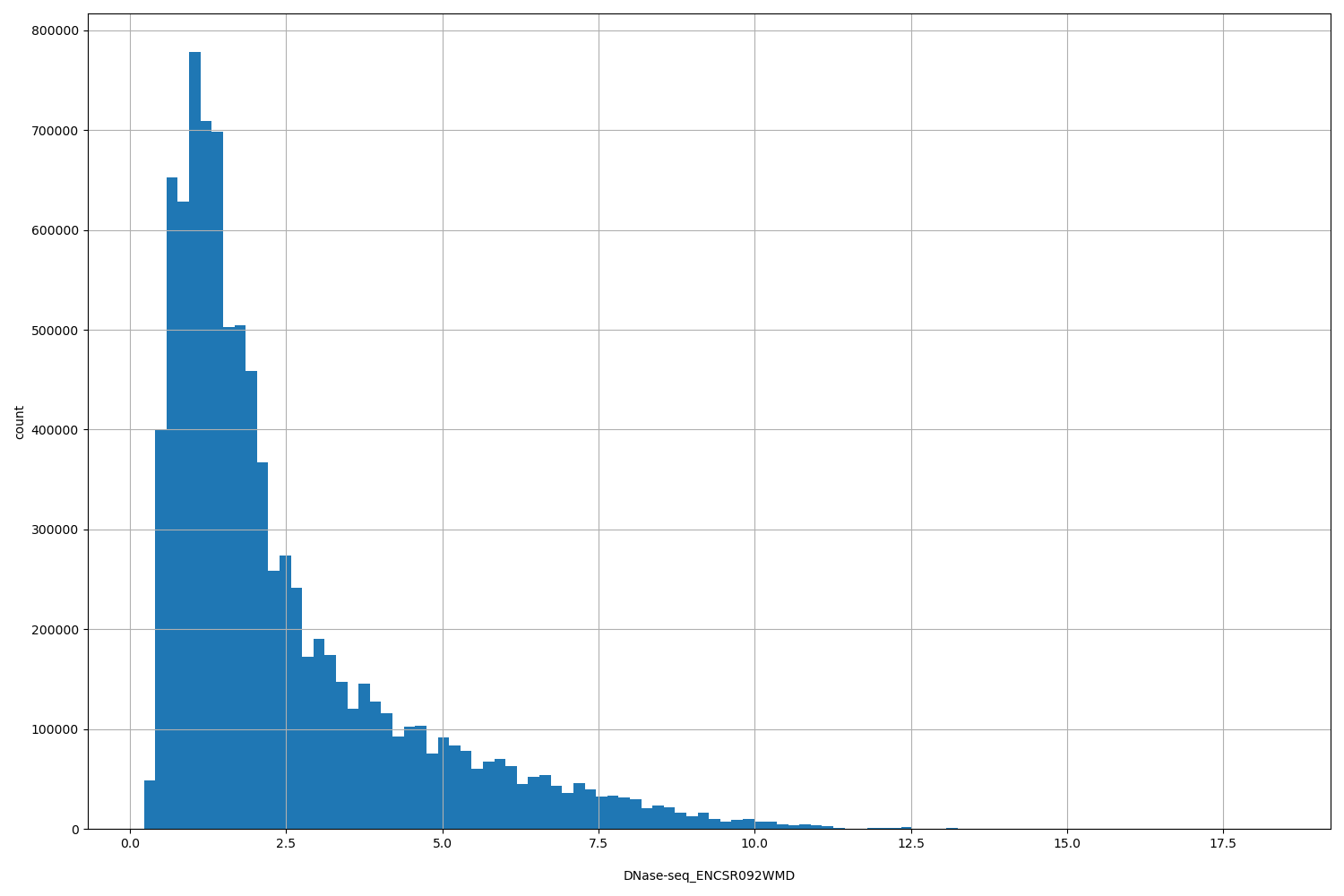
DNase-seq_ENCSR092WMD
| Id: | DNase-seq/ENCSR092WMD |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR092WMD [biosample_summary="Homo sapiens left renal cortex interstitium tissue male embryo (105 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: male embryo (105 days) output_type: peaks audit_warning: Replicate 1_2 has no significant footprints detected. audit_warning: Alignment file {ENCFF394BLL|/files/ENCFF394BLL/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 31752245 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR092WMD | float |
DNase-seq_ENCSR092WMD |
DNase-seq ENCSR092WMD [biosample_summary="Homo sapiens left renal cortex interstitium tissue male embryo (105 days)"]
|
 |
[0.221, 18.3] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF192WAG.bed.gz | 484.45 KB | d638f2b75d6e32a853e415ea5f56687b |
| ENCFF192WAG.bed.gz.dvc | 100.0 B | 4d64c726c65f42bfe45b9d105c505dff |
| ENCFF192WAG.tabix.bed.gz | 437.94 KB | a9cf9e5e360b36104ca81e317f8e834a |
| ENCFF192WAG.tabix.bed.gz.dvc | 106.0 B | 2b840cf02dbd11b69915e9dae79554a8 |
| ENCFF192WAG.tabix.bed.gz.tbi | 251.46 KB | 9f4b086aa9779f76b0d4f954437066f8 |
| ENCFF192WAG.tabix.bed.gz.tbi.dvc | 110.0 B | f42dfe59633b345eeed297634838fc4d |
| genomic_resource.yaml | 1.97 KB | d21df788db7964408ed71cb846cebdc3 |
| genomic_resource_original.yaml | 1.85 KB | 2eeb033c94e4f74e8868ebcbf2ca32aa |
| statistics/ |