DNase-seq_ENCSR083FBK
| Id: | DNase-seq/ENCSR083FBK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR083FBK [biosample_summary="Homo sapiens small intestine tissue female embryo (110 days)"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female embryo (110 days) output_type: peaks audit_internal_action: Archived analysis {ENCAN592UBL|/analyses/ENCAN592UBL/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF697AKY|/files/ENCFF697AKY/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 36488096 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF204MVL|/files/ENCFF204MVL/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 36224662 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
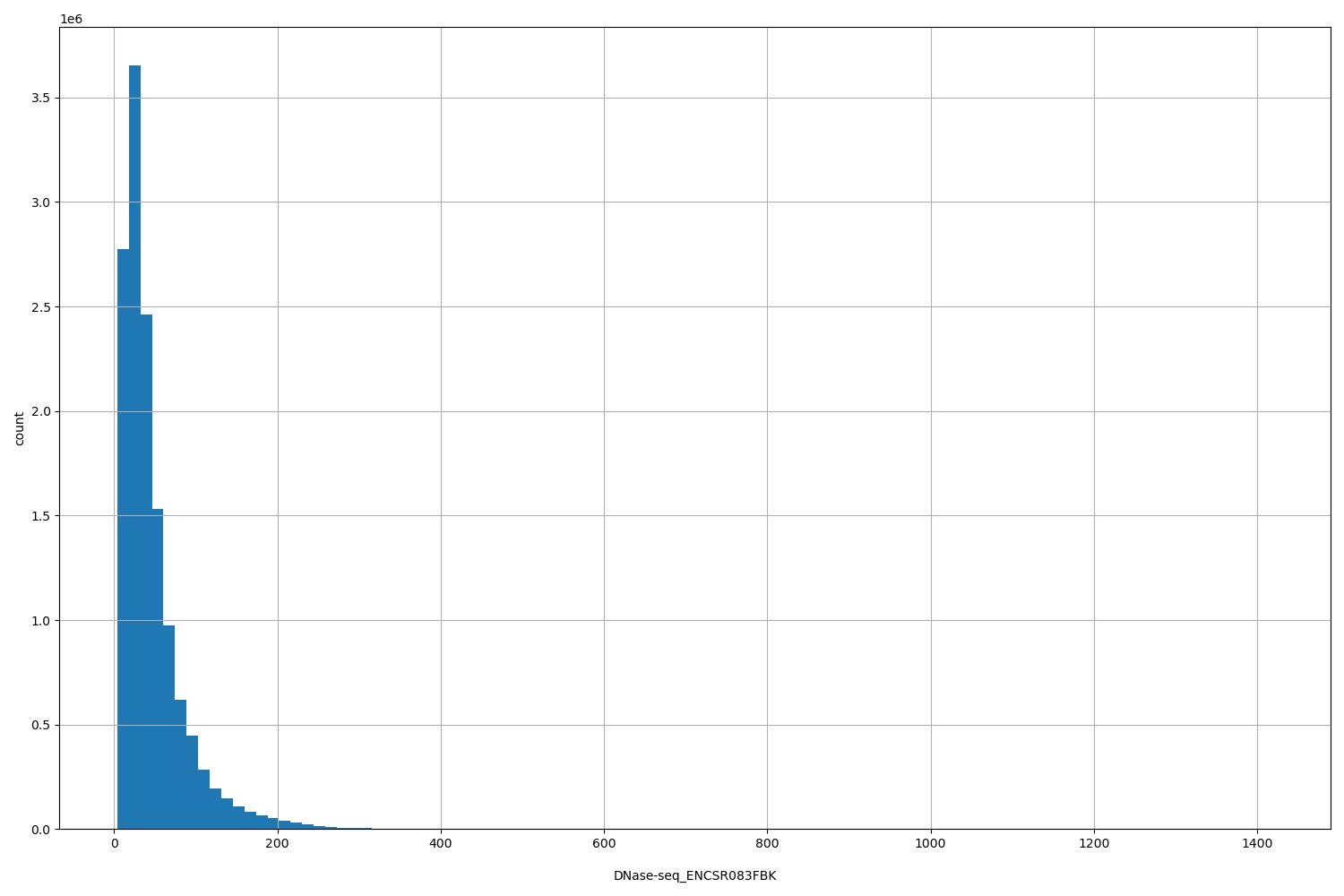
| DNase-seq_ENCSR083FBK | float |
DNase-seq_ENCSR083FBK |
DNase-seq ENCSR083FBK [biosample_summary="Homo sapiens small intestine tissue female embryo (110 days)"]
|
 |
[4, 1.42e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF130CZB.bed.gz | 871.45 KB | 124749e83dfdcecc0b703295613704f0 |
| ENCFF130CZB.bed.gz.dvc | 100.0 B | c74c49a2d3fcce4f644de50ea35c9d4a |
| ENCFF130CZB.tabix.bed.gz | 746.12 KB | c2d8656fe0473e0d436e8ef043060434 |
| ENCFF130CZB.tabix.bed.gz.dvc | 106.0 B | c9734de4f8262318aa72340b95017d4d |
| ENCFF130CZB.tabix.bed.gz.tbi | 422.18 KB | beb8a9c0ee43551b9c06c45dcb2d007f |
| ENCFF130CZB.tabix.bed.gz.tbi.dvc | 110.0 B | 79b3af79345f3094e423cd4fd6559b1e |
| genomic_resource.yaml | 2.61 KB | ed9e94e0bbd28b3a15cdab623acf1fc4 |
| genomic_resource_original.yaml | 2.51 KB | 75211be0e27cf356b3d894a8c381982d |
| statistics/ |