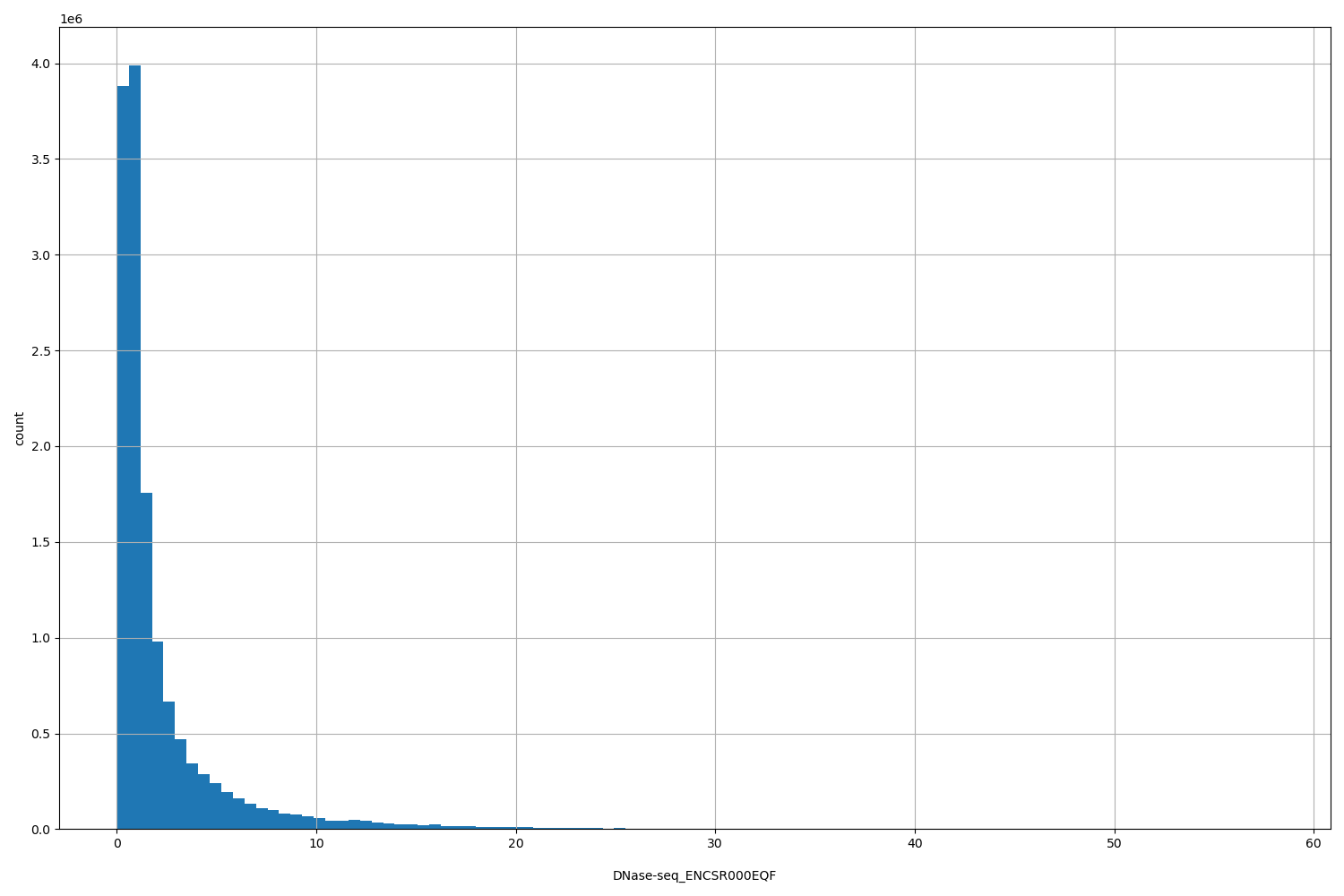
DNase-seq_ENCSR000EQF
| Id: | DNase-seq/ENCSR000EQF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EQF [biosample_summary="Homo sapiens T-helper 17 cell"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Signal Portion of Tags (SPOT) is a measure of enrichment, analogous to the commonly used fraction of reads in peaks metric. ENCODE processed alignment files {ENCFF248FBZ|/files/ENCFF248FBZ/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ({ENCPL848KLD|/pipelines/ENCPL848KLD/}) for GRCh38 assembly have a SPOT1 score of 0.29. According to ENCODE standards, SPOT1 score of 0.4 or higher is considered a product of high quality data. Any sample with a SPOT1 score <0.3 should be targeted for replacement with a higher quality sample, and a SPOT1 score of 0.25 is considered minimally acceptable SPOT1 score of 0.25 is considered minimally acceptable for rare and hard to find primary tissues. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EQF | float |
DNase-seq_ENCSR000EQF |
DNase-seq ENCSR000EQF [biosample_summary="Homo sapiens T-helper 17 cell"]
|
 |
[0.0342, 57.9] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF047UUQ.bed.gz | 832.47 KB | 79c2b996a41269190d45d0cf11f36c78 |
| ENCFF047UUQ.bed.gz.dvc | 100.0 B | 44b68351055adcf85c4fdce40a9838f6 |
| ENCFF047UUQ.tabix.bed.gz | 758.38 KB | 1d121223d8eac015731c0ba57b674ca4 |
| ENCFF047UUQ.tabix.bed.gz.dvc | 106.0 B | 66e53a35b7d26d8efc84f57408e5c436 |
| ENCFF047UUQ.tabix.bed.gz.tbi | 316.2 KB | 1eb417f0d5d868dea431b73825dd1933 |
| ENCFF047UUQ.tabix.bed.gz.tbi.dvc | 110.0 B | 46d5c72ef65fb7c0902ba041e8c8af46 |
| genomic_resource.yaml | 1.95 KB | d67b90cf2483e86ee6147c54b8a1091f |
| genomic_resource_original.yaml | 1.87 KB | 48b986d4617945773ddc316c6df1d26b |
| statistics/ |