DNase-seq_ENCSR000EPM
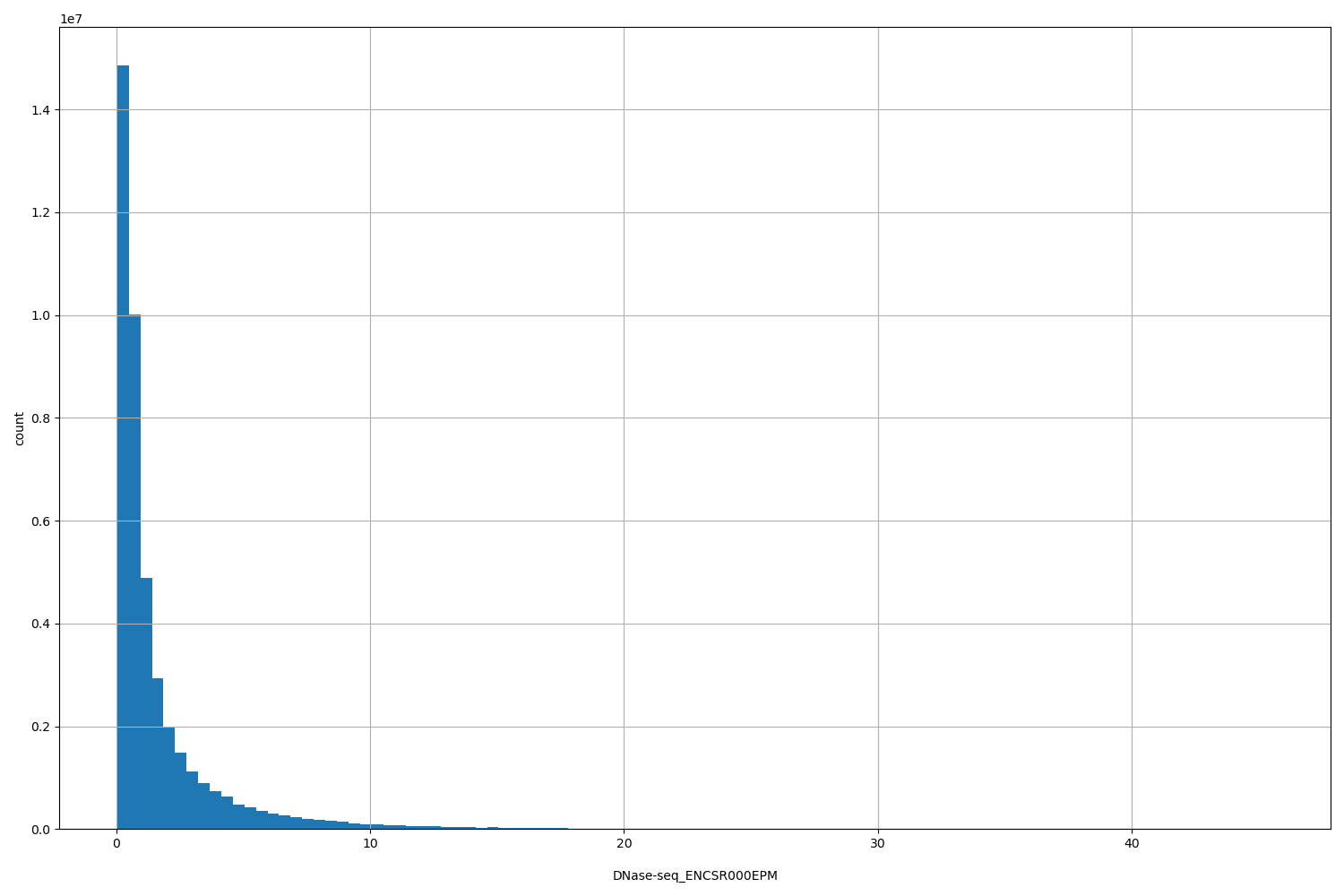
| Id: | DNase-seq/ENCSR000EPM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EPM [biosample_summary="Homo sapiens astrocyte"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Alignment file {ENCFF688FWV|/files/ENCFF688FWV/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 22771395 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EPM | float |
DNase-seq_ENCSR000EPM |
DNase-seq ENCSR000EPM [biosample_summary="Homo sapiens astrocyte"]
|
 |
[0.0409, 45.6] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF503LDV.bed.gz | 2.4 MB | 2cc95f88480a73051447be84cac9f5b1 |
| ENCFF503LDV.bed.gz.dvc | 101.0 B | 619762a7a011e7acd6e535cf50eb11cc |
| ENCFF503LDV.tabix.bed.gz | 2.19 MB | abe78f6922b72e4e2d3776ca068b3246 |
| ENCFF503LDV.tabix.bed.gz.dvc | 107.0 B | d0e1d618af5e1940dc052bd919b77edb |
| ENCFF503LDV.tabix.bed.gz.tbi | 681.57 KB | 9ee136e3e8d70a69430bf2e533d76b2d |
| ENCFF503LDV.tabix.bed.gz.tbi.dvc | 110.0 B | f002e44a93751a9093ef16c64de4d0f2 |
| genomic_resource.yaml | 1.71 KB | 0b64b9136e3a1d16ae8fcd74ec2cdeea |
| genomic_resource_original.yaml | 1.64 KB | 468af3e383fc2e316e1fbc1b1ad41e83 |
| statistics/ |