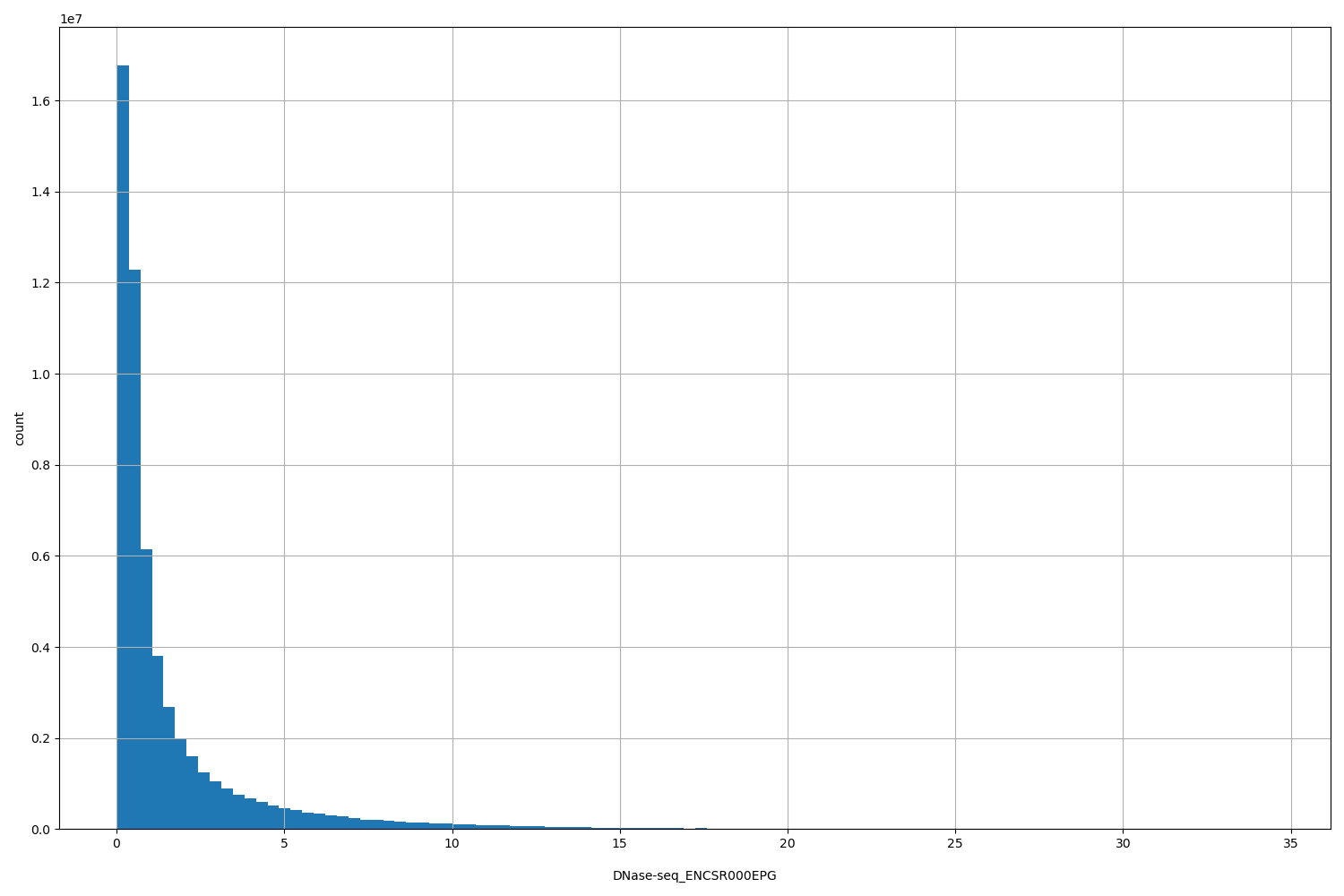
DNase-seq_ENCSR000EPG
| Id: | DNase-seq/ENCSR000EPG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EPG [biosample_summary="Homo sapiens M059J"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Alignment file {ENCFF184FIC|/files/ENCFF184FIC/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 48894458 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EPG | float |
DNase-seq_ENCSR000EPG |
DNase-seq ENCSR000EPG [biosample_summary="Homo sapiens M059J"]
|
 |
[0.0224, 34.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF355POY.bed.gz | 3.13 MB | 1b24a16f70f6df80a378c5cd2a990875 |
| ENCFF355POY.bed.gz.dvc | 101.0 B | 3cb0c8672392628dc8f48d40e6d6fe0e |
| ENCFF355POY.tabix.bed.gz | 2.86 MB | 8482374f230e891eb4ffa1c600755bb2 |
| ENCFF355POY.tabix.bed.gz.dvc | 107.0 B | 8f19444bb0bd4ffb4cc40e5f36431b43 |
| ENCFF355POY.tabix.bed.gz.tbi | 745.47 KB | 14c0ed7a4d9802311f5b2c14bddfe138 |
| ENCFF355POY.tabix.bed.gz.tbi.dvc | 110.0 B | 0d61a0e5ea313221b89551315deecd26 |
| genomic_resource.yaml | 1.7 KB | e3e5139a8450c930968c71d61a7b021b |
| genomic_resource_original.yaml | 1.63 KB | 48378affcceb260c8df209bf3733ad38 |
| statistics/ |