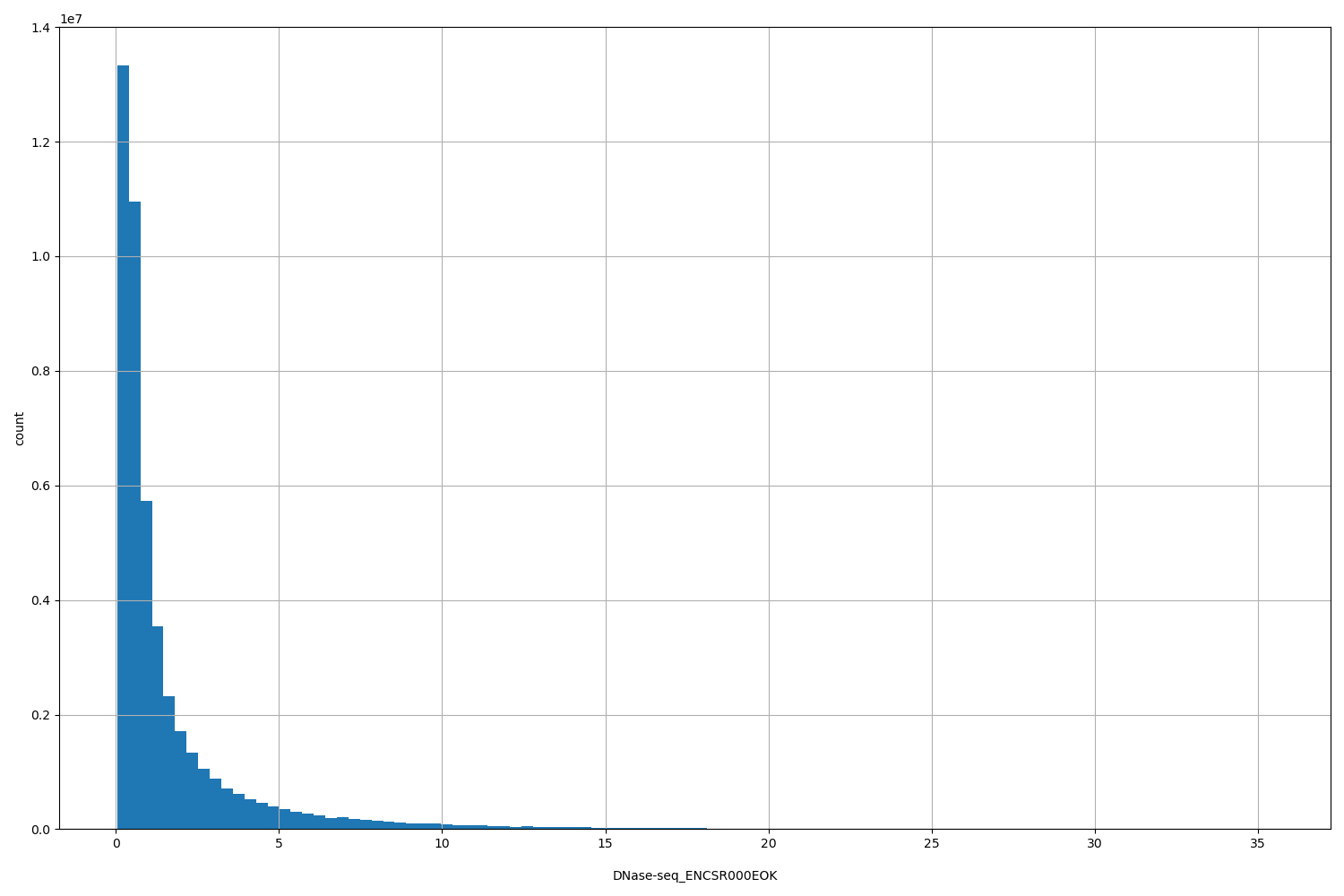
DNase-seq_ENCSR000EOK
| Id: | DNase-seq/ENCSR000EOK |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000EOK [biosample_summary="Homo sapiens renal cortical epithelial cell"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_warning: Alignment file {ENCFF767UJT|/files/ENCFF767UJT/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 22043063 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000EOK | float |
DNase-seq_ENCSR000EOK |
DNase-seq ENCSR000EOK [biosample_summary="Homo sapiens renal cortical epithelial cell"]
|
 |
[0.0467, 35.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF303ROT.bed.gz | 2.67 MB | 55481873efc624adc7da23c9157dadcd |
| ENCFF303ROT.bed.gz.dvc | 101.0 B | 640d3a2e199b3ec527a381553ae50abf |
| ENCFF303ROT.tabix.bed.gz | 2.43 MB | 7d8bf084fed92b9df8ecbfaab7d48f54 |
| ENCFF303ROT.tabix.bed.gz.dvc | 107.0 B | 88da0515e4abed427b7287c4849c1271 |
| ENCFF303ROT.tabix.bed.gz.tbi | 731.31 KB | c90a7c5aed30c10be0fdda3464ec97af |
| ENCFF303ROT.tabix.bed.gz.tbi.dvc | 110.0 B | 6bf7c834e84a32653a05cb954ae17a12 |
| genomic_resource.yaml | 1.77 KB | e6a54ab38d9dcc2571c31563dd356136 |
| genomic_resource_original.yaml | 1.68 KB | 247d80204f612e2bba5dc2fb858795e6 |
| statistics/ |