DNase-seq_ENCSR000ENY
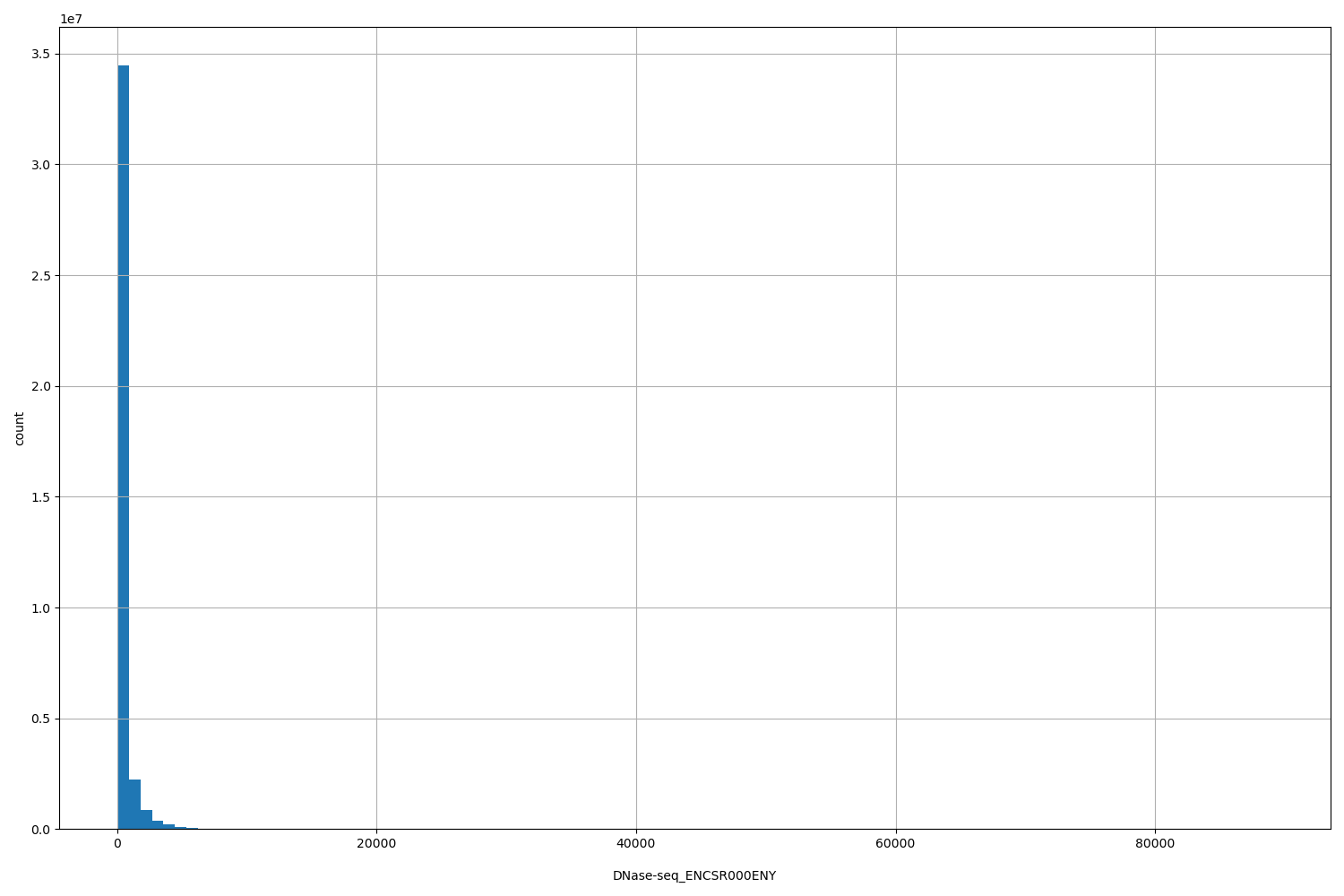
| Id: | DNase-seq/ENCSR000ENY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000ENY [biosample_summary="Homo sapiens dermis blood vessel endothelial cell female adult"] |
| Description: |
status: released biological_replicates: Rep 1 summary: female adult output_type: peaks audit_internal_action: Archived analysis {ENCAN669FHR|/analyses/ENCAN669FHR/} has in progress subobject quality standard {encode3-dnase|/quality-standards/encode3-dnase/} audit_warning: Alignment file {ENCFF490NJA|/files/ENCFF490NJA/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 45684134 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) audit_warning: Alignment file {ENCFF200WYL|/files/ENCFF200WYL/} processed by DNase-seq ENCODE3 GRCh38 pipeline ( {ENCPL001DNS|/pipelines/ENCPL001DNS/} ) for GRCh38 assembly has 45469809 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq/}) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000ENY | float |
DNase-seq_ENCSR000ENY |
DNase-seq ENCSR000ENY [biosample_summary="Homo sapiens dermis blood vessel endothelial cell female adult"]
|
 |
[1, 8.91e+04] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF894ZLD.bed.gz | 2.43 MB | 7b02a7e4136c561bebd5c34c48648a4a |
| ENCFF894ZLD.bed.gz.dvc | 101.0 B | e01482b38a745ca7c200b5262603c031 |
| ENCFF894ZLD.tabix.bed.gz | 2.11 MB | f059870014bc8a6435039c5987a74dc6 |
| ENCFF894ZLD.tabix.bed.gz.dvc | 107.0 B | 2e58197d801540caac42dd499e5c2fd2 |
| ENCFF894ZLD.tabix.bed.gz.tbi | 702.41 KB | 023bc9ceed080d85c98c5b84ba7c26a5 |
| ENCFF894ZLD.tabix.bed.gz.tbi.dvc | 110.0 B | b95aa2c1856292a49f4c1489c229d76d |
| genomic_resource.yaml | 2.59 KB | 58d920d6d1fbebbd98c7f205e335d306 |
| genomic_resource_original.yaml | 2.49 KB | 6ddf40403570200121820a7bbd034bec |
| statistics/ |