DNase-seq_ENCSR000ENR
| Id: | DNase-seq/ENCSR000ENR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
DNase-seq ENCSR000ENR [biosample_summary="Homo sapiens HFF-Myc originated from foreskin fibroblast"] |
| Description: |
status: released biological_replicates: Rep 1 summary: output_type: peaks audit_error: Alignment file {ENCFF992SGG|/files/ENCFF992SGG/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 19550895 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) audit_warning: Alignment file {ENCFF988KRZ|/files/ENCFF988KRZ/} processed by DNase-seq ENCODE4 v3.0.0-alpha.2 GRCh38 pipeline ( {ENCPL848KLD|/pipelines/ENCPL848KLD/} ) for GRCh38 assembly has 29170145 mapped reads. According to ENCODE standards, conventional DNase-seq profile requires a minimum of 20 million uniquely mapped reads to generate a reliable SPOT (Signal Portion of Tags) score. The recommended value is > 50 million. For deep, foot-printing depth DNase-seq 150-200 million uniquely mapped reads are recommended. (See {ENCODE DNase-seq data standards|/data-standards/dnase-seq-encode4/}) |
| Labels: |
|
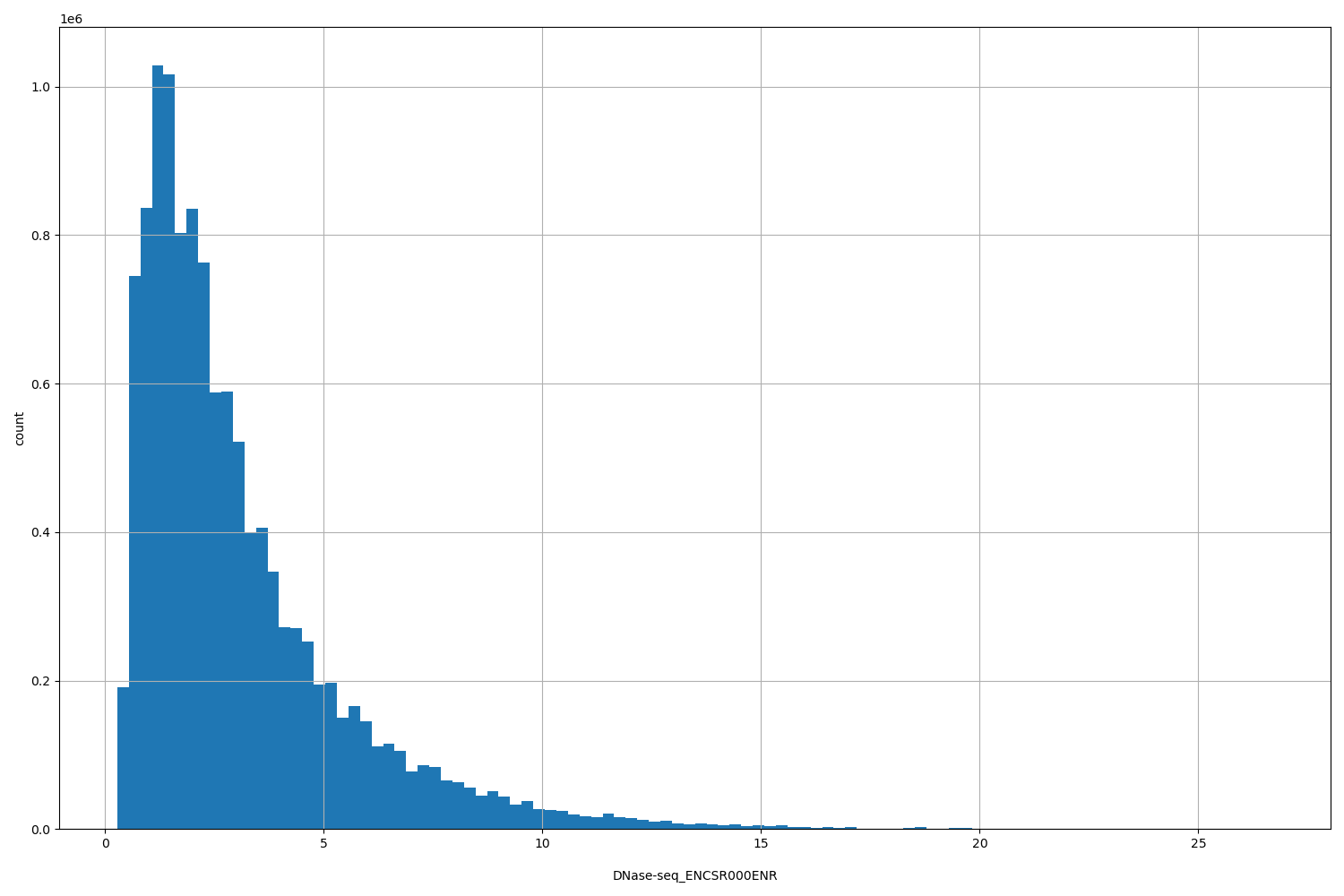
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| DNase-seq_ENCSR000ENR | float |
DNase-seq_ENCSR000ENR |
DNase-seq ENCSR000ENR [biosample_summary="Homo sapiens HFF-Myc originated from foreskin fibroblast"]
|
 |
[0.276, 26.7] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF030SHD.bed.gz | 709.39 KB | e15cf8cfdeb4c38951fd3f4cae8189cd |
| ENCFF030SHD.bed.gz.dvc | 100.0 B | 7cd3bbe2ffe3b59fb0aaf3b698cb6cce |
| ENCFF030SHD.tabix.bed.gz | 628.05 KB | 0c46b4af605d1d9e4b6fbcd44eb5f206 |
| ENCFF030SHD.tabix.bed.gz.dvc | 106.0 B | 25a42a04b895291c9982b100e7f4c8e5 |
| ENCFF030SHD.tabix.bed.gz.tbi | 367.46 KB | cef74f5add364da10e38767f1450c423 |
| ENCFF030SHD.tabix.bed.gz.tbi.dvc | 110.0 B | f47a6569c163c00a01cd8bfc8a53789c |
| genomic_resource.yaml | 2.45 KB | b4a5458572fb2f101976ddd425930e39 |
| genomic_resource_original.yaml | 2.35 KB | ef3fa84fcdffc330ec7cf261eb18ab66 |
| statistics/ |